The expression platform and the aptamer: cooperativity between Mg2+ and ligand in the SAM-I riboswitch
- PMID: 23258703
- PMCID: PMC3562059
- DOI: 10.1093/nar/gks978
The expression platform and the aptamer: cooperativity between Mg2+ and ligand in the SAM-I riboswitch
Abstract
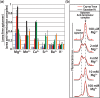
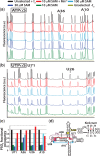
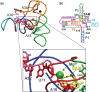
Riboswitch operation involves the complex interplay between the aptamer domain and the expression platform. During transcription, these two domains compete against each other for shared sequence. In this study, we explore the cooperative effects of ligand binding and Magnesium interactions in the SAM-I riboswitch in the context of aptamer collapse and anti-terminator formation. Overall, our studies show the apo-aptamer acts as (i) a pre-organized aptamer competent to bind ligand and undergo structural collapse and (ii) a conformation that is more accessible to anti-terminator formation. We show that both Mg(2+) ions and SAM are required for a collapse transition to occur. We then use competition between the aptamer and expression platform for shared sequence to characterize the stability of the collapsed aptamer. We find that SAM and Mg(2+) interactions in the aptamer are highly cooperative in maintaining switch polarity (i.e. aptamer 'off-state' versus anti-terminator 'on-state'). We further show that the aptamer off-state is preferentially stabilized by Mg(2+) and similar divalent ions. Furthermore, the functional switching assay was used to select for phosphorothioate interference, and identifies potential magnesium chelation sites while characterizing their coordinated role with SAM in aptamer stabilization. In addition, we find that Mg(2+) interactions with the apo-aptamer are required for the full formation of the anti-terminator structure, and that higher concentrations of Mg(2+) (>4 mM) shift the equilibrium toward the anti-terminator on-state even in the presence of SAM.
Figures






Similar articles
-
Magnesium controls aptamer-expression platform switching in the SAM-I riboswitch.Nucleic Acids Res. 2019 Apr 8;47(6):3158-3170. doi: 10.1093/nar/gky1311. Nucleic Acids Res. 2019. PMID: 30605518 Free PMC article.
-
Cooperation between Magnesium and Metabolite Controls Collapse of the SAM-I Riboswitch.Biophys J. 2017 Jul 25;113(2):348-359. doi: 10.1016/j.bpj.2017.06.044. Biophys J. 2017. PMID: 28746845 Free PMC article.
-
Tertiary contacts control switching of the SAM-I riboswitch.Nucleic Acids Res. 2011 Mar;39(6):2416-31. doi: 10.1093/nar/gkq1096. Epub 2010 Nov 19. Nucleic Acids Res. 2011. PMID: 21097777 Free PMC article.
-
Structure-based insights into recognition and regulation of SAM-sensing riboswitches.Sci China Life Sci. 2023 Jan;66(1):31-50. doi: 10.1007/s11427-022-2188-7. Epub 2022 Nov 29. Sci China Life Sci. 2023. PMID: 36459353 Review.
-
Riboswitches that sense S-adenosylmethionine and S-adenosylhomocysteine.Biochem Cell Biol. 2008 Apr;86(2):157-68. doi: 10.1139/O08-008. Biochem Cell Biol. 2008. PMID: 18443629 Review.
Cited by
-
Getting to the bottom of lncRNA mechanism: structure-function relationships.Mamm Genome. 2022 Jun;33(2):343-353. doi: 10.1007/s00335-021-09924-x. Epub 2021 Oct 12. Mamm Genome. 2022. PMID: 34642784 Free PMC article. Review.
-
Potential effects of metal ion induced two-state allostery on the regulatory mechanism of add adenine riboswitch.Commun Biol. 2022 Oct 22;5(1):1120. doi: 10.1038/s42003-022-04096-z. Commun Biol. 2022. PMID: 36273041 Free PMC article.
-
Magnesium controls aptamer-expression platform switching in the SAM-I riboswitch.Nucleic Acids Res. 2019 Apr 8;47(6):3158-3170. doi: 10.1093/nar/gky1311. Nucleic Acids Res. 2019. PMID: 30605518 Free PMC article.
-
Magnesium ions mediate ligand binding and conformational transition of the SAM/SAH riboswitch.Commun Biol. 2023 Jul 31;6(1):791. doi: 10.1038/s42003-023-05175-5. Commun Biol. 2023. PMID: 37524918 Free PMC article.
-
Common themes and differences in SAM recognition among SAM riboswitches.Biochim Biophys Acta. 2014 Oct;1839(10):931-938. doi: 10.1016/j.bbagrm.2014.05.013. Epub 2014 May 23. Biochim Biophys Acta. 2014. PMID: 24863160 Free PMC article. Review.
References
-
- Mandal M, Breaker RR. Adenine riboswitches and gene activation by disruption of a transcription terminator. Nat. Struct. Mol. Biol. 2004;11:29–35. - PubMed
-
- Winkler WC, Nahvi A, Sudarsan N, Barrick JE, Breaker RR. An mRNA structure that controls gene expression by binding S-adenosylmethionine. Nat. Struct. Biol. 2003;10:701–707. - PubMed
-
- Mandal M, Lee M, Barrick JE, Weinberg Z, Emilsson GM, Ruzzo WL, Breaker RR. A glycine-dependent riboswitch that uses cooperative binding to control gene expression. Science. 2004;306:275–279. - PubMed

