Structural and mechanistic studies of HpxO, a novel flavin adenine dinucleotide-dependent urate oxidase from Klebsiella pneumoniae
- PMID: 23259842
- PMCID: PMC3552057
- DOI: 10.1021/bi301262p
Structural and mechanistic studies of HpxO, a novel flavin adenine dinucleotide-dependent urate oxidase from Klebsiella pneumoniae
Abstract
HpxO is a flavin-dependent urate oxidase that catalyzes the hydroxylation of uric acid to 5-hydroxyisourate and functions in a novel pathway for purine catabolism found in Klebsiella pneumoniae. We have determined the structures of HpxO with and without uric acid at 2.0 and 2.2 Å, respectively. We have also determined the structure of the R204Q variant at 2.0 Å resolution in the absence of uric acid. The variant structure is very similar to that of wild-type HpxO except for the conformation of Arg103, which interacts with FAD in the variant but not in the wild-type structure. Interestingly, the R204Q variant results in the uncoupling of nicotinamide adenine dinucleotide oxidation from uric acid hydroxylation. This suggests that Arg204 facilitates the deprotonation of uric acid, activating it for the oxygen transfer. On the basis of these data, a mechanism for this reaction consisting of a nucleophilic attack of the urate anion on the flavin hydroperoxide resulting in the formation of 5-hydroxyisourate is proposed.
Figures


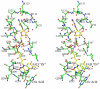


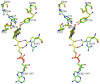
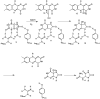
References
-
- Vanwynsberghe T, Verhamme K, Raymaekers M, Cartuyvels R, Van Vaerenbergh K, Boel A, de Beenhouwer H. A Large Hospital Outbreak of Klebsiella pneumoniae (DHA-1 and SHV-11 Positive): Importance of Detection and Treatment of ampC-Lactamases. The Open Infectious Diseases J. 2009;3:55–60.
-
- Ko W-C, Paterson DL, Sagnimeni AJ, Hansen DS, Von Gottberg A, Mohapatra S, Casellas JM, Goossens H, Mulazimoglu L, Trenholme G, Klugman KP, McCormack JG, Yu VL. Community-Acquired Klebsiella pneumoniae Bacteremia: Global Differences in Clinical Patterns. Emerging Infectious Diseases. 2002;8:160–166. - PMC - PubMed
-
- Maltezou HC. Metallo-β-lactamases in Gram-negative bacteria: introducing the era of pan-resistance? Int. J. Antimicrobial Agents. 2009;β33:405.e401–405.e407. - PubMed
-
- Nordmann P, Cuzon G, Naas T. The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. The Lancet. 2009;9:228–236. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

