miR156 and miR390 regulate tasiRNA accumulation and developmental timing in Physcomitrella patens
- PMID: 23263766
- PMCID: PMC3556961
- DOI: 10.1105/tpc.112.103176
miR156 and miR390 regulate tasiRNA accumulation and developmental timing in Physcomitrella patens
Abstract
microRNA156 (miR156) affects developmental timing in flowering plants. miR156 and its target relationships with members of the SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) gene family appear universally conserved in land plants, but the specific functions of miR156 outside of flowering plants are unknown. We find that miR156 promotes a developmental change from young filamentous protonemata to leafy gametophores in the moss Physcomitrella patens, opposite to its role as an inhibitor of development in flowering plants. P. patens miR156 also influences accumulation of trans-acting small interfering RNAs (tasiRNAs) dependent upon a second ancient microRNA, miR390. Both miR156 and miR390 directly target a single major tasiRNA primary transcript. Inhibition of miR156 function causes increased miR390-triggered tasiRNA accumulation and decreased accumulation of tasiRNA targets. Overexpression of miR390 also caused a slower formation of gametophores, elevated miR390-triggered tasiRNA accumulation, and reduced level of tasiRNA targets. We conclude that a gene regulatory network controlled by miR156, miR390, and their targets controls developmental change in P. patens. The broad outlines and regulatory logic of this network are conserved in flowering plants, albeit with some modifications. Partially conserved small RNA networks thus influence developmental timing in plants with radically different body plans.
Figures






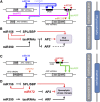
References
-
- Adenot X., Elmayan T., Lauressergues D., Boutet S., Bouché N., Gasciolli V., Vaucheret H. (2006). DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr. Biol. 16: 927–932 - PubMed
-
- Arazi T., Talmor-Neiman M., Stav R., Riese M., Huijser P., Baulcombe D.C. (2005). Cloning and characterization of micro-RNAs from moss. Plant J. 43: 837–848 - PubMed
-
- Ashton N.W., Cove D.J. (1977). The isolation and preliminary characterization of auxotropic and analogue-resistant mutants of the moss Physcomitrella patens. Mol. Gen. Genet. 154: 87–95
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

