Feeding and the rhodopsin family g-protein coupled receptors in nematodes and arthropods
- PMID: 23264768
- PMCID: PMC3524798
- DOI: 10.3389/fendo.2012.00157
Feeding and the rhodopsin family g-protein coupled receptors in nematodes and arthropods
Abstract
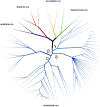
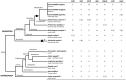
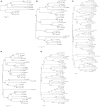
In vertebrates, receptors of the rhodopsin G-protein coupled superfamily (GPCRs) play an important role in the regulation of feeding and energy homeostasis and are activated by peptide hormones produced in the brain-gut axis. These peptides regulate appetite and energy expenditure by promoting or inhibiting food intake. Sequence and function homologs of human GPCRs involved in feeding exist in the nematode roundworm, Caenorhabditis elegans (C. elegans), and the arthropod fruit fly, Drosophila melanogaster (D. melanogaster), suggesting that the mechanisms that regulate food intake emerged early and have been conserved during metazoan radiation. Nematodes and arthropods are the most diverse and successful animal phyla on Earth. They can survive in a vast diversity of environments and have acquired distinct life styles and feeding strategies. The aim of the present review is to investigate if this diversity has affected the evolution of invertebrate GPCRs. Homologs of the C. elegans and D. melanogaster rhodopsin receptors were characterized in the genome of other nematodes and arthropods and receptor evolution compared. With the exception of bombesin receptors (BBR) that are absent from nematodes, a similar gene complement was found. In arthropods, rhodopsin GPCR evolution is characterized by species-specific gene duplications and deletions and in nematodes by gene expansions in species with a free-living stage and gene deletions in representatives of obligate parasitic taxa. Based upon variation in GPCR gene number and potentially divergent functions within phyla we hypothesize that life style and feeding diversity practiced by nematodes and arthropods was one factor that contributed to rhodopsin GPCR gene evolution. Understanding how the regulation of food intake has evolved in invertebrates will contribute to the development of novel drugs to control nematodes and arthropods and the pests and diseases that use them as vectors.
Keywords: conservation; evolution; feeding; invertebrates; rhodopsin GPCR.
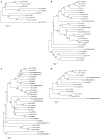
Figures


Similar articles
-
Nematode and arthropod genomes provide new insights into the evolution of class 2 B1 GPCRs.PLoS One. 2014 Mar 20;9(3):e92220. doi: 10.1371/journal.pone.0092220. eCollection 2014. PLoS One. 2014. PMID: 24651821 Free PMC article.
-
Evolution of secretin family GPCR members in the metazoa.BMC Evol Biol. 2006 Dec 13;6:108. doi: 10.1186/1471-2148-6-108. BMC Evol Biol. 2006. PMID: 17166275 Free PMC article.
-
Prevertebrate Local Gene Duplication Facilitated Expansion of the Neuropeptide GPCR Superfamily.Mol Biol Evol. 2015 Nov;32(11):2803-17. doi: 10.1093/molbev/msv179. Epub 2015 Sep 3. Mol Biol Evol. 2015. PMID: 26337547
-
A genome-wide inventory of neurohormone GPCRs in the red flour beetle Tribolium castaneum.Front Neuroendocrinol. 2008 Jan;29(1):142-65. doi: 10.1016/j.yfrne.2007.10.003. Epub 2007 Oct 24. Front Neuroendocrinol. 2008. PMID: 18054377 Review.
-
Allatostatin-type A, kisspeptin and galanin GPCRs and putative ligands as candidate regulatory factors of mantle function.Mar Genomics. 2016 Jun;27:25-35. doi: 10.1016/j.margen.2015.12.003. Epub 2015 Dec 30. Mar Genomics. 2016. PMID: 26751715 Review.
Cited by
-
Evolution and Potential Function in Molluscs of Neuropeptide and Receptor Homologues of the Insect Allatostatins.Front Endocrinol (Lausanne). 2021 Sep 29;12:725022. doi: 10.3389/fendo.2021.725022. eCollection 2021. Front Endocrinol (Lausanne). 2021. PMID: 34659116 Free PMC article.
-
Palmitoylation of Prolactin-Releasing Peptide Increased Affinity for and Activation of the GPR10, NPFF-R2 and NPFF-R1 Receptors: In Vitro Study.Int J Mol Sci. 2021 Aug 18;22(16):8904. doi: 10.3390/ijms22168904. Int J Mol Sci. 2021. PMID: 34445614 Free PMC article.
-
Phylogenetic investigation of Peptide hormone and growth factor receptors in five dipteran genomes.Front Endocrinol (Lausanne). 2013 Dec 16;4:193. doi: 10.3389/fendo.2013.00193. eCollection 2013. Front Endocrinol (Lausanne). 2013. PMID: 24379806 Free PMC article.
-
Receptors Mediating Host-Microbiota Communication in the Metaorganism: The Invertebrate Perspective.Front Immunol. 2020 Jun 16;11:1251. doi: 10.3389/fimmu.2020.01251. eCollection 2020. Front Immunol. 2020. PMID: 32612612 Free PMC article. Review.
-
Global analysis of neuropeptide receptor conservation across phylum Nematoda.BMC Biol. 2024 Oct 8;22(1):223. doi: 10.1186/s12915-024-02017-6. BMC Biol. 2024. PMID: 39379997 Free PMC article.
References
-
- Aguilar R., Maestro J. L., Vilaplana L., Pascual N., Piulachs M. D., Belles X. (2003). Allatostatin gene expression in brain and midgut, and activity of synthetic allatostatins on feeding-related processes in the cockroach Blattella germanica. Regul. Pept. 115, 171–17710.1016/S0167-0115(03)00165-4 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

