Recruitment of Arf1-GDP to Golgi by Glo3p-type ArfGAPs is crucial for golgi maintenance and plant growth
- PMID: 23266962
- PMCID: PMC3561012
- DOI: 10.1104/pp.112.209148
Recruitment of Arf1-GDP to Golgi by Glo3p-type ArfGAPs is crucial for golgi maintenance and plant growth
Abstract
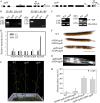
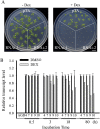
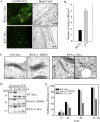
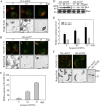
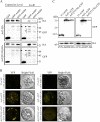
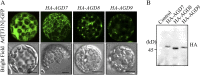
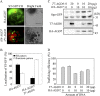
ADP-ribosylation factor1 (Arf1), a member of the small GTP-binding proteins, plays a pivotal role in protein trafficking to multiple organelles. In its GDP-bound form, Arf1 is recruited from the cytosol to organelle membranes, where it functions in vesicle-mediated protein trafficking. However, the mechanism of Arf1-GDP recruitment remains unknown. Here, we provide evidence that two Glo3p-type Arf GTPase-activating proteins (ArfGAPs), ArfGAP domain8 (AGD8) and AGD9, are involved in the recruitment of Arf1-GDP to the Golgi apparatus in Arabidopsis (Arabidopsis thaliana). RNA interference plants expressing low levels of AGD8 and AGD9 exhibited abnormal Golgi morphology, inhibition of protein trafficking, and arrest of plant growth and development. In RNA interference plants, Arf1 was poorly recruited to the Golgi apparatus. Conversely, high levels of AGD8 and AGD9 induced Arf1 accumulation at the Golgi and suppressed Golgi disruption and inhibition of vacuolar trafficking that was caused by overexpression of AGD7. Based on these results, we propose that the Glo3p-type ArfGAPs AGD8 and AGD9 recruit Arf1-GDP from the cytosol to the Golgi for Arf1-mediated protein trafficking, which is essential for plant development and growth.
Figures


References
-
- Abramoff MD, Magelhaes PJ, Ram SJ. (2004) Image processing with ImageJ. Biophotonics Int 11: 36–42
-
- Aoyama T, Chua NH. (1997) A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J 11: 605–612 - PubMed
-
- Balch WE, Kahn RA, Schwaninger R. (1992) ADP-ribosylation factor is required for vesicular trafficking between the endoplasmic reticulum and the cis-Golgi compartment. J Biol Chem 267: 13053–13061 - PubMed
-
- Bigay J, Gounon P, Robineau S, Antonny B. (2003) Lipid packing sensed by ArfGAP1 couples COPI coat disassembly to membrane bilayer curvature. Nature 426: 563–566 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

