Reannotation of translational start sites in the genome of Mycobacterium tuberculosis
- PMID: 23273318
- PMCID: PMC3582765
- DOI: 10.1016/j.tube.2012.11.012
Reannotation of translational start sites in the genome of Mycobacterium tuberculosis
Abstract
Identification and correction of incorrect ORF start sites is important for a variety of experimental and analytical purposes, ranging from cloning to inference of operon structure. The genome of the H37Rv reference strain of Mycobacterium tuberculosis (Mtb) was originally annotated when it was first sequenced nearly 15 years ago. While this annotation has served the TB research community well as a standard of reference for over a decade, it has been demonstrated experimentally that the actual start sites for an estimated 5-10% of open reading frames differ from the annotation. In this paper, we present a comprehensive bioinformatic analysis of all 3989 ORFs (open reading frames) in the M. tuberculosis H37Rv genome. Our method combines information from comparative analysis (alignment to start sites of orthologs in other Actinobacteria), sequence conservation, "protein likeness", putative ribosome binding sites, and other data to identify translational start sites. The features are combined in a linear model that is trained on dataset of known start sites verified by mass spectrometry, with a cross-validated accuracy of 94%. The method can be viewed as an augmentation of Hidden Markov Model-based tools such as Glimmer and GeneMark by incorporating more information than just the raw genomic sequence to decide which position is the legitimate translational start site for each ORF. Using this analysis, we identify 269 genes that most likely need to be re-annotated, and identify the best alterative translational start site for each. These revised ORF definitions could be used in the reannotation of the H37Rv genome, as well as to prioritize genes for experimental start-site validation.
Copyright © 2012 Elsevier Ltd. All rights reserved.
Figures
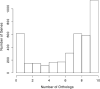
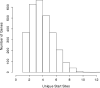
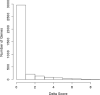

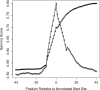
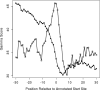
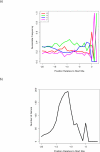
Similar articles
-
Proteogenomic analysis of polymorphisms and gene annotation divergences in prokaryotes using a clustered mass spectrometry-friendly database.Mol Cell Proteomics. 2011 Jan;10(1):M110.002527. doi: 10.1074/mcp.M110.002527. Epub 2010 Oct 28. Mol Cell Proteomics. 2011. PMID: 21030493 Free PMC article.
-
Experimental determination of translational start sites resolves uncertainties in genomic open reading frame predictions - application to Mycobacterium tuberculosis.Microbiology (Reading). 2009 Jan;155(Pt 1):186-197. doi: 10.1099/mic.0.022889-0. Microbiology (Reading). 2009. PMID: 19118359 Free PMC article.
-
[Analysis, identification and correction of some errors of model refseqs appeared in NCBI Human Gene Database by in silico cloning and experimental verification of novel human genes].Yi Chuan Xue Bao. 2004 May;31(5):431-43. Yi Chuan Xue Bao. 2004. PMID: 15478601 Chinese.
-
Non-AUG start codons: Expanding and regulating the small and alternative ORFeome.Exp Cell Res. 2020 Jun 1;391(1):111973. doi: 10.1016/j.yexcr.2020.111973. Epub 2020 Mar 21. Exp Cell Res. 2020. PMID: 32209305 Free PMC article. Review.
-
Updating and curating metabolic pathways of TB.Tuberculosis (Edinb). 2013 Jan;93(1):47-59. doi: 10.1016/j.tube.2012.11.001. Epub 2013 Feb 1. Tuberculosis (Edinb). 2013. PMID: 23375378 Free PMC article. Review.
Cited by
-
Structural and functional insight into the Mycobacterium tuberculosis protein PrpR reveals a novel type of transcription factor.Nucleic Acids Res. 2019 Oct 10;47(18):9934-9949. doi: 10.1093/nar/gkz724. Nucleic Acids Res. 2019. PMID: 31504787 Free PMC article.
-
Construction of an overexpression library for Mycobacterium tuberculosis.Biol Methods Protoc. 2018;3(1):bpy009. doi: 10.1093/biomethods/bpy009. Epub 2018 Aug 20. Biol Methods Protoc. 2018. PMID: 30197930 Free PMC article.
-
Genome-wide mapping of transcriptional start sites defines an extensive leaderless transcriptome in Mycobacterium tuberculosis.Cell Rep. 2013 Nov 27;5(4):1121-31. doi: 10.1016/j.celrep.2013.10.031. Epub 2013 Nov 21. Cell Rep. 2013. PMID: 24268774 Free PMC article.
-
Environmental Sensing in Actinobacteria: a Comprehensive Survey on the Signaling Capacity of This Phylum.J Bacteriol. 2015 Aug 1;197(15):2517-35. doi: 10.1128/JB.00176-15. Epub 2015 May 18. J Bacteriol. 2015. PMID: 25986905 Free PMC article.
-
A novel regulatory interplay between atypical B12 riboswitches and uORF translation in Mycobacterium tuberculosis.Nucleic Acids Res. 2024 Jul 22;52(13):7876-7892. doi: 10.1093/nar/gkae338. Nucleic Acids Res. 2024. PMID: 38709884 Free PMC article.
References
-
- Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon SV, Eiglmeier K, Gas S, Barry CE, 3rd, Tekaia F, Badcock K, Basham D, Brown D, Chillingworth T, Connor R, Davies R, Devlin K, Feltwell T, Gentles S, Hamlin N, Holroyd S, Hornsby T, Jagels K, Krogh A, McLean J, Moule S, Murphy L, Oliver K, Osborne J, Quail MA, Rajandream MA, Rogers J, Rutter S, Seeger K, Skelton J, Squares R, Squares S, Sulston JE, Taylor K, Whitehead S, Barrell BG. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature. 1998;393:537–544. doi: 10.1038/31159. - PubMed
-
- Camus JC, Pryor MJ, Medigue C, Cole ST. Re-annotation of the genome sequence of Mycobacterium tuberculosis H37Rv. Microbiology. 2002;148:2967–2973. - PubMed
-
- Smollett KL, Fivian-Hughes AS, Smith JE, Chang A, Rao T, Davis EO. Experimental determination of translational start sites resolves uncertainties in genomic open reading frame predictions - application to Mycobacterium tuberculosis. Microbiology. 2009;155:186–197. doi: 155/1/186 [pii] 10.1099/mic.0.022889-0. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

