Meta-analysis followed by replication identifies loci in or near CDKN1B, TET3, CD80, DRAM1, and ARID5B as associated with systemic lupus erythematosus in Asians
- PMID: 23273568
- PMCID: PMC3542470
- DOI: 10.1016/j.ajhg.2012.11.018
Meta-analysis followed by replication identifies loci in or near CDKN1B, TET3, CD80, DRAM1, and ARID5B as associated with systemic lupus erythematosus in Asians
Abstract
Systemic lupus erythematosus (SLE) is a prototype autoimmune disease with a strong genetic involvement and ethnic differences. Susceptibility genes identified so far only explain a small portion of the genetic heritability of SLE, suggesting that many more loci are yet to be uncovered for this disease. In this study, we performed a meta-analysis of genome-wide association studies on SLE in Chinese Han populations and followed up the findings by replication in four additional Asian cohorts with a total of 5,365 cases and 10,054 corresponding controls. We identified genetic variants in or near CDKN1B, TET3, CD80, DRAM1, and ARID5B as associated with the disease. These findings point to potential roles of cell-cycle regulation, autophagy, and DNA demethylation in SLE pathogenesis. For the region involving TET3 and that involving CDKN1B, multiple independent SNPs were identified, highlighting a phenomenon that might partially explain the missing heritability of complex diseases.
Copyright © 2013 The American Society of Human Genetics. Published by Elsevier Inc. All rights reserved.
Figures

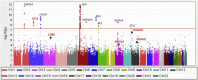



References
-
- Arnett F.C., Shulman L.E. Studies in familial systemic lupus erythematosus. Medicine (Baltimore) 1976;55:313–322. - PubMed
-
- Mok C.C., Lau C.S. Lupus in Hong Kong Chinese. Lupus. 2003;12:717–722. - PubMed
-
- Yang W., Ng P., Zhao M., Hirankarn N., Lau C.S., Mok C.C., Chan T.M., Wong R.W., Lee K.W., Mok M.Y. Population differences in SLE susceptibility genes: STAT4 and BLK, but not PXK, are associated with systemic lupus erythematosus in Hong Kong Chinese. Genes Immun. 2009;10:219–226. - PubMed
-
- Yang W., Shen N., Ye D.Q., Liu Q., Zhang Y., Qian X.X., Hirankarn N., Ying D., Pan H.F., Mok C.C., Asian Lupus Genetics Consortium Genome-wide association study in Asian populations identifies variants in ETS1 and WDFY4 associated with systemic lupus erythematosus. PLoS Genet. 2010;6:e1000841. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

