Comparative genome analysis of six malarial parasites using codon usage bias based tools
- PMID: 23275725
- PMCID: PMC3530877
- DOI: 10.6026/97320630081230
Comparative genome analysis of six malarial parasites using codon usage bias based tools
Abstract
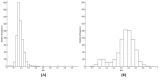
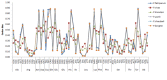
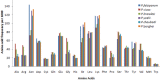
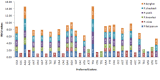
Codon usage bias (CUB) is an omnipresent phenomenon, which occurs in nearly all organisms. Previous studies of codon bias in Plasmodium species were based on a limited dataset. This study uses whole genome datasets for comparative genome analysis of six Plasmodium species using CUB and other related methods for the first time. Codon usage bias, compositional variation in translated amino acid frequency, effective number of codons and optimal codons are analyzed for P.falciparum, P.vivax, P.knowlesi, P.berghei, P.chabaudii and P.yoelli. A plot of effective number of codons versus GC3 shows their differential codon usage pattern arises due to a combination of mutational and translational selection pressure. The increased relative usage of adenine and thymine ending optimal codons in highly expressed genes of P.falciparum is the result of higher composition biased pressure, and usage of guanine and cytosine bases at third codon position can be explained by translational selection pressure acting on them. While higher usage of adenine and thymine bases at third codon position in optimal codons of P.vivax highlights the role of translational selection pressure apart from composition biased mutation pressure in shaping their codon usage pattern. The frequency of those amino acids that are encoded by AT ending codons are significantly high in P.falciparum due to action of high composition biased mutational pressure compared with other Plasmodium species. The CUB variation in the three rodent parasites, P.berghei, P.chabaudii and P.yoelli is strikingly similar to that of P.falciparum. The simian and human malarial parasite, P.knowlesi shows a variation in codon usage bias similar to P.vivax but on closer study there are differences confirmed by the method of Principal Component Analysis (PCA).
Abbreviations: CDS - Coding sequences, GC1 - GC composition at first site of codon, GC2 - GC composition at second site of codon, GC3 - GC composition at third site of codon, Ala - Alanine, Arg - Arginine, Asn - Asparagine, Asp - Aspartic acid, Cys - Cysteine, Gln - Glutamine Glu - Glutamic acid Gly - Glycine His - Histidine Ile - Isoleucine Leu - Leucine Lys - Lysine Met - Methionine Phe - Phenylalanine Pro - Proline Ser - Serine Thr - Threonine Trp - Tryptophan Tyr - Tyrosine Val - Valine.
Keywords: Codon usage bias (CUB); Effective number of codon (ENC); Optimal codon; Principal Component Analysis (PCA); RSCU; tRNA.
Figures


Similar articles
-
Comparative analysis of codon usage patterns of Plasmodium helical interspersed subtelomeric (PHIST) proteins.Front Microbiol. 2023 Dec 14;14:1320060. doi: 10.3389/fmicb.2023.1320060. eCollection 2023. Front Microbiol. 2023. PMID: 38156001 Free PMC article.
-
Analysis of codon usage patterns in Haloxylon ammodendron based on genomic and transcriptomic data.Gene. 2022 Dec 15;845:146842. doi: 10.1016/j.gene.2022.146842. Epub 2022 Aug 28. Gene. 2022. PMID: 36038027
-
Comparison of codon usage and tRNAs in mitochondrial genomes of Candida species.Biosystems. 2007 Sep-Oct;90(2):362-70. doi: 10.1016/j.biosystems.2006.09.039. Epub 2006 Oct 5. Biosystems. 2007. PMID: 17123703
-
Analysis of codon usage bias of thioredoxin in apicomplexan protozoa.Parasit Vectors. 2023 Nov 21;16(1):431. doi: 10.1186/s13071-023-06002-w. Parasit Vectors. 2023. PMID: 37990340 Free PMC article.
-
New insights into the factors affecting synonymous codon usage in human infecting Plasmodium species.Acta Trop. 2017 Dec;176:29-33. doi: 10.1016/j.actatropica.2017.07.025. Epub 2017 Jul 24. Acta Trop. 2017. PMID: 28751162 Review.
Cited by
-
Asparagine requirement in Plasmodium berghei as a target to prevent malaria transmission and liver infections.Nat Commun. 2015 Nov 4;6:8775. doi: 10.1038/ncomms9775. Nat Commun. 2015. PMID: 26531182 Free PMC article.
-
Comparative Analysis of the Codon Usage Pattern in the Chloroplast Genomes of Gnetales Species.Int J Mol Sci. 2024 Oct 2;25(19):10622. doi: 10.3390/ijms251910622. Int J Mol Sci. 2024. PMID: 39408952 Free PMC article.
-
Cryptosporidium felis differs from other Cryptosporidium spp. in codon usage.Microb Genom. 2021 Dec;7(12):000711. doi: 10.1099/mgen.0.000711. Microb Genom. 2021. PMID: 34907893 Free PMC article.
-
Evolutionary pressures and codon bias in low complexity regions of plasmodia.Genetica. 2021 Aug;149(4):217-237. doi: 10.1007/s10709-021-00126-6. Epub 2021 Jul 12. Genetica. 2021. PMID: 34254217
-
Mitochondrial genomes of Macropsini (Hemiptera: Cicadellidae: Eurymelinae): Structural features, codon usage patterns, and phylogenetic implications.Ecol Evol. 2024 Sep 10;14(9):e70268. doi: 10.1002/ece3.70268. eCollection 2024 Sep. Ecol Evol. 2024. PMID: 39263460 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
