Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection
- PMID: 23277552
- PMCID: PMC3549110
- DOI: 10.1073/pnas.1214550110
Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection
Abstract
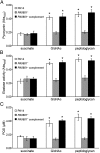
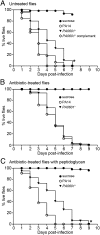
Most infections result from colonization by more than one microbe. Within such polymicrobial infections, microbes often display synergistic interactions that result in increased disease severity. Although many clinical studies have documented the occurrence of synergy in polymicrobial infections, little is known about the underlying molecular mechanisms. A prominent pathogen in many polymicrobial infections is Pseudomonas aeruginosa, a Gram-negative bacterium that displays enhanced virulence during coculture with Gram-positive bacteria. In this study we discovered that during coinfection, P. aeruginosa uses peptidoglycan shed by Gram-positive bacteria as a cue to stimulate production of multiple extracellular factors that possess lytic activity against prokaryotic and eukaryotic cells. Consequently, P. aeruginosa displays enhanced virulence in a Drosophila model of infection when cocultured with Gram-positive bacteria. Inactivation of a gene (PA0601) required for peptidoglycan sensing mitigated this phenotype. Using Drosophila and murine models of infection, we also show that peptidoglycan sensing results in P. aeruginosa-mediated reduction in the Gram-positive flora in the infection site. Our data suggest that P. aeruginosa has evolved a mechanism to survey the microbial community and respond to Gram-positive produced peptidoglycan through production of antimicrobials and toxins that not only modify the composition of the community but also enhance host killing. Additionally, our results suggest that therapeutic strategies targeting Gram-positive bacteria might be a viable approach for reducing the severity of P. aeruginosa polymicrobial infections.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Pasteur L, Joubert J. Charbon et septicemie. Compt Rend Acad. 1877;85:101–105.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

