Mouse prion protein polymorphism Phe-108/Val-189 affects the kinetics of fibril formation and the response to seeding: evidence for a two-step nucleation polymerization mechanism
- PMID: 23283973
- PMCID: PMC3576082
- DOI: 10.1074/jbc.M112.414581
Mouse prion protein polymorphism Phe-108/Val-189 affects the kinetics of fibril formation and the response to seeding: evidence for a two-step nucleation polymerization mechanism
Abstract
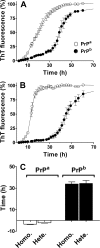
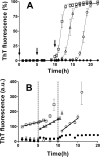
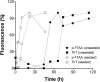
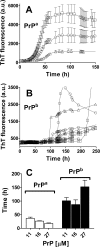
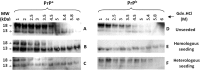
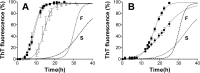
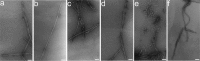
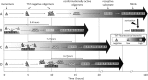
Prion diseases are fatal neurodegenerative disorders associated with the polymerization of the cellular form of prion protein (PrP(C)) into an amyloidogenic β-sheet infectious form (PrP(Sc)). The sequence of host PrP is the major determinant of host prion disease susceptibility. In mice, the presence of allele a (Prnp(a), encoding the polymorphism Leu-108/Thr-189) or b (Prnp(b), Phe-108/Val-189) is associated with short or long incubation times, respectively, following infection with PrP(Sc). The molecular bases linking PrP sequence, infection susceptibility, and convertibility of PrP(C) into PrP(Sc) remain unclear. Here we show that recombinant PrP(a) and PrP(b) aggregate and respond to seeding differently in vitro. Our kinetic studies reveal differences during the nucleation phase of the aggregation process, where PrP(b) exhibits a longer lag phase that cannot be completely eliminated by seeding the reaction with preformed fibrils. Additionally, PrP(b) is more prone to propagate features of the seeds, as demonstrated by conformational stability and electron microscopy studies of the formed fibrils. We propose a model of polymerization to explain how the polymorphisms at positions 108 and 189 produce the phenotypes seen in vivo. This model also provides insight into phenomena such as species barrier and prion strain generation, two phenomena also influenced by the primary structure of PrP.
Figures

Comment in
-
Implications of prion polymorphisms.Prion. 2013 Jul-Aug;7(4):276-9. doi: 10.4161/pri.25566. Epub 2013 Jun 27. Prion. 2013. PMID: 23807178 Free PMC article. Review.
Similar articles
-
Implications of prion polymorphisms.Prion. 2013 Jul-Aug;7(4):276-9. doi: 10.4161/pri.25566. Epub 2013 Jun 27. Prion. 2013. PMID: 23807178 Free PMC article. Review.
-
Role of N-terminal familial mutations in prion protein fibrillization and prion amyloid propagation in vitro.J Biol Chem. 2006 Mar 24;281(12):8190-6. doi: 10.1074/jbc.M513417200. Epub 2006 Jan 26. J Biol Chem. 2006. PMID: 16443601
-
Soluble polymorphic bank vole prion proteins induced by co-expression of quiescin sulfhydryl oxidase in E. coli and their aggregation behaviors.Microb Cell Fact. 2017 Oct 4;16(1):170. doi: 10.1186/s12934-017-0782-x. Microb Cell Fact. 2017. PMID: 28978309 Free PMC article.
-
Autocatalytic conversion of recombinant prion proteins displays a species barrier.J Biol Chem. 2004 Feb 27;279(9):7671-7. doi: 10.1074/jbc.M310594200. Epub 2003 Dec 10. J Biol Chem. 2004. PMID: 14668351
-
Prion protein and its conformational conversion: a structural perspective.Top Curr Chem. 2011;305:135-67. doi: 10.1007/128_2011_165. Top Curr Chem. 2011. PMID: 21630136 Review.
Cited by
-
Pharmacological chaperone reshapes the energy landscape for folding and aggregation of the prion protein.Nat Commun. 2016 Jun 27;7:12058. doi: 10.1038/ncomms12058. Nat Commun. 2016. PMID: 27346148 Free PMC article.
-
Probing the origin of prion protein misfolding via reconstruction of ancestral proteins.Protein Sci. 2022 Dec;31(12):e4477. doi: 10.1002/pro.4477. Protein Sci. 2022. PMID: 36254680 Free PMC article.
-
Implications of prion polymorphisms.Prion. 2013 Jul-Aug;7(4):276-9. doi: 10.4161/pri.25566. Epub 2013 Jun 27. Prion. 2013. PMID: 23807178 Free PMC article. Review.
-
Beta-endoproteolysis of the cellular prion protein by dipeptidyl peptidase-4 and fibroblast activation protein.Proc Natl Acad Sci U S A. 2023 Jan 3;120(1):e2209815120. doi: 10.1073/pnas.2209815120. Epub 2022 Dec 27. Proc Natl Acad Sci U S A. 2023. PMID: 36574660 Free PMC article.
-
The role of the unusual threonine string in the conversion of prion protein.Sci Rep. 2016 Dec 16;6:38877. doi: 10.1038/srep38877. Sci Rep. 2016. PMID: 27982059 Free PMC article.
References
-
- Prusiner S. B. (1982) Novel proteinaceous infectious particles cause scrapie. Science 216, 136–144 - PubMed
-
- Bruce M. E., Fraser H. (1991) Scrapie strain variation and its implications. Curr. Top. Microbiol. Immunol. 172, 125–138 - PubMed
-
- Fraser H., Dickinson A. G. (1973) Scrapie in mice. Agent-strain differences in the distribution and intensity of grey matter vacuolation. J. Comp. Pathol. 83, 29–40 - PubMed
-
- Bruce M. E. (1993) Scrapie strain variation and mutation. Br. Med. Bull. 49, 822–838 - PubMed
-
- Gibbs C. J., Jr., Gajdusek D. C. (1973) Experimental subacute spongiform virus encephalopathies in primates and other laboratory animals. Science 182, 67–68 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

