Translocation and stability of replicative DNA helicases upon encountering DNA-protein cross-links
- PMID: 23283980
- PMCID: PMC3576070
- DOI: 10.1074/jbc.M112.419358
Translocation and stability of replicative DNA helicases upon encountering DNA-protein cross-links
Abstract
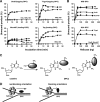
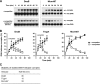
DNA-protein cross-links (DPCs) are formed when cells are exposed to various DNA-damaging agents. Because DPCs are extremely large, steric hindrance conferred by DPCs is likely to affect many aspects of DNA transactions. In DNA replication, DPCs are first encountered by the replicative helicase that moves at the head of the replisome. However, little is known about how replicative helicases respond to covalently immobilized protein roadblocks. In the present study we elucidated the effect of DPCs on the DNA unwinding reaction of hexameric replicative helicases in vitro using defined DPC substrates. DPCs on the translocating strand but not on the nontranslocating strand impeded the progression of the helicases including the phage T7 gene 4 protein, simian virus 40 large T antigen, Escherichia coli DnaB protein, and human minichromosome maintenance Mcm467 subcomplex. The impediment varied with the size of the cross-linked proteins, with a threshold size for clearance of 5.0-14.1 kDa. These results indicate that the central channel of the dynamically translocating hexameric ring helicases can accommodate only small proteins and that all of the helicases tested use the steric exclusion mechanism to unwind duplex DNA. These results further suggest that DPCs on the translocating and nontranslocating strands constitute helicase and polymerase blocks, respectively. The helicases stalled by DPC had limited stability and dissociated from DNA with a half-life of 15-36 min. The implications of the results are discussed in relation to the distinct stabilities of replisomes that encounter tight but reversible DNA-protein complexes and irreversible DPC roadblocks.
Figures




References
-
- Johnson A., O'Donnell M. (2005) Cellular DNA replicases: components and dynamics at the replication fork. Annu. Rev. Biochem. 74, 283–315 - PubMed
-
- Hamdan S. M., Richardson C. C. (2009) Motors, switches, and contacts in the replisome. Annu. Rev. Biochem. 78, 205–243 - PubMed
-
- Alabert C., Groth A. (2012) Chromatin replication and epigenome maintenance. Nat. Rev. Mol. Cell Biol. 13, 153–167 - PubMed
-
- Masai H., Matsumoto S., You Z., Yoshizawa-Sugata N., Oda M. (2010) Eukaryotic chromosome DNA replication: where, when, and how? Annu. Rev. Biochem. 79, 89–130 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

