Replication of Epstein-Barr viral DNA
- PMID: 23284049
- PMCID: PMC3579399
- DOI: 10.1101/cshperspect.a013029
Replication of Epstein-Barr viral DNA
Abstract
Epstein-Barr virus (EBV) is a paradigm for human tumor viruses: it is the first virus recognized to cause cancer in people; it causes both lymphomas and carcinomas; yet these tumors arise infrequently given that most people in the world are infected with the virus. EBV is maintained extrachromosomally in infected normal and tumor cells. Eighty-four percent of these viral plasmids replicate each S phase, are licensed, require a single viral protein for their synthesis, and can use two functionally distinct origins of DNA replication, oriP, and Raji ori. Eighty-eight percent of newly synthesized plasmids are segregated faithfully to the daughter cells. Infectious viral particles are not synthesized under these conditions of latent infection. This plasmid replication is consistent with survival of EBV's host cells. Rare cells in an infected population either spontaneously or following exogenous induction support EBV's lytic cycle, which is lethal for the cell. In this case, the viral DNA replicates 100-fold or more, uses a third kind of viral origin of DNA replication, oriLyt, and many viral proteins. Here we shall describe the three modes of EBV's replication as a function of the viral origins used and the viral and cellular proteins that mediate the DNA synthesis from these origins focusing, where practical, on recent advances in our understanding.
Figures
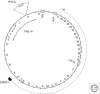
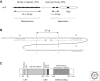
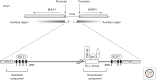
References
-
- Antson AA, Burns JE, Moroz OV, Scott DJ, Sanders CM, Bronstein IB, Dodson GG, Wilson KS, Maitland NJ 2000. Structure of the intact transactivation domain of the human papillomavirus E2 protein. Nature 403: 805–809 - PubMed
-
- Avolio-Hunter TM, Frappier L 2003. EBNA1 efficiently assembles on chromatin containing the Epstein–Barr virus latent origin of replication. Virology 315: 398–408 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
