N-methyl-D-aspartate receptor-dependent denitrosylation of neuronal nitric oxide synthase increase the enzyme activity
- PMID: 23285183
- PMCID: PMC3532120
- DOI: 10.1371/journal.pone.0052788
N-methyl-D-aspartate receptor-dependent denitrosylation of neuronal nitric oxide synthase increase the enzyme activity
Abstract
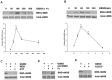
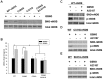
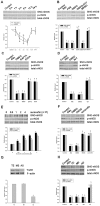
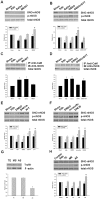
Our laboratory once reported that neuronal nitric oxide synthase (nNOS) S-nitrosylation was decreased in rat hippocampus during cerebral ischemia-reperfusion, but the underlying mechanism was unclear. In this study, we show that nNOS activity is dynamically regulated by S-nitrosylation. We found that overexpressed nNOS in HEK293 (human embryonic kidney) cells could be S-nitrosylated by exogenous NO donor GSNO and which is associated with the enzyme activity decrease. Cys(331), one of the zinc-tetrathiolate cysteines, was identified as the key site of nNOS S-nitrosylation. In addition, we also found that nNOS is highly S-nitrosylated in resting rat hippocampal neurons and the enzyme undergos denitrosylation during the process of rat brain ischemia/reperfusion. Intrestingly, the process of nNOS denitrosylation is coupling with the decrease of nNOS phosphorylation at Ser(847), a site associated with nNOS activation. Further more, we document that nNOS denitrosylation could be suppressed by pretreatment of neurons with MK801, an antagonist of NMDAR, GSNO, EGTA, BAPTA, W-7, an inhibitor of calmodulin as well as TrxR1 antisense oligonucleotide (AS-ODN) respectively. Taken together, our data demonstrate that the denitrosylation of nNOS induced by calcium ion influx is a NMDAR-dependent process during the early stage of ischemia/reperfusion, which is majorly mediated by thioredoxin-1 (Trx1) system. nNOS dephosphorylation may be induced by the enzyme denitrosylation, which suggest that S-nitrosylation/denitrosylation of nNOS may be an important mechanism in regulating the enzyme activity.
Conflict of interest statement
Figures




Similar articles
-
Denitrosylation of nNOS induced by cerebral ischemia-reperfusion contributes to nitrosylation of CaMKII and its inhibition of autophosphorylation in hippocampal CA1.Eur Rev Med Pharmacol Sci. 2019 Sep;23(17):7674-7683. doi: 10.26355/eurrev_201909_18891. Eur Rev Med Pharmacol Sci. 2019. PMID: 31539160
-
S-nitrosylation of c-Src via NMDAR-nNOS module promotes c-Src activation and NR2A phosphorylation in cerebral ischemia/reperfusion.Mol Cell Biochem. 2012 Jun;365(1-2):363-77. doi: 10.1007/s11010-012-1280-4. Epub 2012 Mar 16. Mol Cell Biochem. 2012. PMID: 22422045
-
Endogenous nitric oxide induces activation of apoptosis signal-regulating kinase 1 via S-nitrosylation in rat hippocampus during cerebral ischemia-reperfusion.Neuroscience. 2013 Jan 15;229:36-48. doi: 10.1016/j.neuroscience.2012.10.055. Epub 2012 Nov 5. Neuroscience. 2013. PMID: 23137546
-
Coordination between Calcium/Calmodulin-Dependent Protein Kinase II and Neuronal Nitric Oxide Synthase in Neurons.Int J Mol Sci. 2020 Oct 27;21(21):7997. doi: 10.3390/ijms21217997. Int J Mol Sci. 2020. PMID: 33121174 Free PMC article. Review.
-
Targeting neuronal nitric oxide synthase and the nitrergic system in post-traumatic stress disorder.Psychopharmacology (Berl). 2022 Oct;239(10):3057-3082. doi: 10.1007/s00213-022-06212-7. Epub 2022 Aug 27. Psychopharmacology (Berl). 2022. PMID: 36029333 Review.
Cited by
-
Procaspase-9 induces its cleavage by transnitrosylating XIAP via the Thioredoxin system during cerebral ischemia-reperfusion in rats.Sci Rep. 2016 Apr 7;6:24203. doi: 10.1038/srep24203. Sci Rep. 2016. PMID: 27052476 Free PMC article.
-
Blocking a vicious cycle nNOS/peroxynitrite/AMPK by S-nitrosoglutathione: implication for stroke therapy.BMC Neurosci. 2015 Jul 15;16:42. doi: 10.1186/s12868-015-0179-x. BMC Neurosci. 2015. PMID: 26174015 Free PMC article.
-
Selective Irreversible Inhibition of Neuronal and Inducible Nitric-oxide Synthase in the Combined Presence of Hydrogen Sulfide and Nitric Oxide.J Biol Chem. 2015 Oct 9;290(41):24932-44. doi: 10.1074/jbc.M115.660316. Epub 2015 Aug 20. J Biol Chem. 2015. PMID: 26296888 Free PMC article.
-
Nitrergic modulation of ion channel function in regulating neuronal excitability.Channels (Austin). 2021 Dec;15(1):666-679. doi: 10.1080/19336950.2021.2002594. Channels (Austin). 2021. PMID: 34802368 Free PMC article.
-
S-nitrosylation of NOS pathway mediators in the penis contributes to cavernous nerve injury-induced erectile dysfunction.Int J Impot Res. 2018 Jun;30(3):108-116. doi: 10.1038/s41443-018-0021-y. Epub 2018 May 8. Int J Impot Res. 2018. PMID: 29736011 Free PMC article.
References
-
- Andrew PJ, Mayer B (1999) Enzymatic function of nitric oxide synthases. Cardiovasc Res 43: 521–531. - PubMed
-
- Jaffrey SR (2005) Detection and characterization of protein nitrosothiols. Methods in Enzymology 396: 105–118. - PubMed
-
- López-Sánchez LM, Corrales FJ, De La Mata M, Muntané J, Rodríguez-Ariza A (2008) Detection and proteomic identification of S-nitrosated proteins in human hepatocytes. Methods Enzymol 440: 273–281. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

