Zinc drives a tertiary fold in the prion protein with familial disease mutation sites at the interface
- PMID: 23290724
- PMCID: PMC3570608
- DOI: 10.1016/j.str.2012.12.002
Zinc drives a tertiary fold in the prion protein with familial disease mutation sites at the interface
Abstract
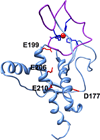
The cellular prion protein PrP(C) consists of two domains--a flexible N-terminal domain, which participates in copper and zinc regulation, and a largely helical C-terminal domain that converts to β sheet in the course of prion disease. These two domains are thought to be fully independent and noninteracting. Compelling cellular and biophysical studies, however, suggest a higher order structure that is relevant to both PrP(C) function and misfolding in disease. Here, we identify a Zn²⁺-driven N-terminal to C-terminal tertiary interaction in PrP(C). The C-terminal surface participating in this interaction carries the majority of the point mutations that confer familial prion disease. Investigation of mutant PrPs finds a systematic relationship between the type of mutation and the apparent strength of this domain structure. The structural features identified here suggest mechanisms by which physiologic metal ions trigger PrP(C) trafficking and control prion disease.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures







References
-
- Aguzzi A, Baumann F, Bremer J. The prion's elusive reason for being. Annu Rev Neurosci. 2008;31:439–477. - PubMed
-
- Bellingham SA, Coleman LA, Masters CL, Camakaris J, Hill AF. Regulation of prion gene expression by transcription factors SP1 and metal transcription factor-1. J Biol Chem. 2009;284:1291–1301. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

