Dissecting the in vivo assembly of the 30S ribosomal subunit reveals the role of RimM and general features of the assembly process
- PMID: 23293003
- PMCID: PMC3575805
- DOI: 10.1093/nar/gks1256
Dissecting the in vivo assembly of the 30S ribosomal subunit reveals the role of RimM and general features of the assembly process
Abstract
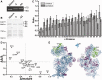
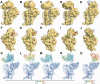
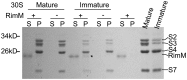
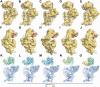
Ribosome biogenesis is a tightly regulated, multi-stepped process. The assembly of ribosomal subunits is a central step of the complex biogenesis process, involving nearly 30 protein factors in vivo in bacteria. Although the assembly process has been extensively studied in vitro for over 40 years, very limited information is known for the in vivo process and specific roles of assembly factors. Such an example is ribosome maturation factor M (RimM), a factor involved in the late-stage assembly of the 30S subunit. Here, we combined quantitative mass spectrometry and cryo-electron microscopy to characterize the in vivo 30S assembly intermediates isolated from mutant Escherichia coli strains with genes for assembly factors deleted. Our compositional and structural data show that the assembly of the 3'-domain of the 30S subunit is severely delayed in these intermediates, featured with highly underrepresented 3'-domain proteins and large conformational difference compared with the mature 30S subunit. Further analysis indicates that RimM functions not only to promote the assembly of a few 3'-domain proteins but also to stabilize the rRNA tertiary structure. More importantly, this study reveals intriguing similarities and dissimilarities between the in vitro and the in vivo assembly pathways, suggesting that they are in general similar but with subtle differences.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

