SRC-2 coactivator deficiency decreases functional reserve in response to pressure overload of mouse heart
- PMID: 23300926
- PMCID: PMC3534027
- DOI: 10.1371/journal.pone.0053395
SRC-2 coactivator deficiency decreases functional reserve in response to pressure overload of mouse heart
Abstract
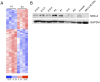
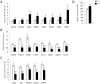
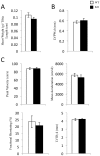
A major component of the cardiac stress response is the simultaneous activation of several gene regulatory networks. Interestingly, the transcriptional regulator steroid receptor coactivator-2, SRC-2 is often decreased during cardiac failure in humans. We postulated that SRC-2 suppression plays a mechanistic role in the stress response and that SRC-2 activity is an important regulator of the adult heart gene expression profile. Genome-wide microarray analysis, confirmed with targeted gene expression analyses revealed that genetic ablation of SRC-2 activates the "fetal gene program" in adult mice as manifested by shifts in expression of a) metabolic and b) sarcomeric genes, as well as associated modulating transcription factors. While these gene expression changes were not accompanied by changes in left ventricular weight or cardiac function, imposition of transverse aortic constriction (TAC) predisposed SRC-2 knockout (KO) mice to stress-induced cardiac dysfunction. In addition, SRC-2 KO mice lacked the normal ventricular hypertrophic response as indicated through heart weight, left ventricular wall thickness, and blunted molecular signaling known to activate hypertrophy. Our results indicate that SRC-2 is involved in maintenance of the steady-state adult heart transcriptional profile, with its ablation inducing transcriptional changes that mimic a stressed heart. These results further suggest that SRC-2 deletion interferes with the timing and integration needed to respond efficiently to stress through disruption of metabolic and sarcomeric gene expression and hypertrophic signaling, the three key stress responsive pathways.
Conflict of interest statement
Figures




References
-
- Asakura M, Kitakaze M (2009) Global gene expression profiling in the failing myocardium. Circulation journal : official journal of the Japanese Circulation Society 73: 1568–1576. - PubMed
-
- Barth AS, Kuner R, Buness A, Ruschhaupt M, Merk S, et al. (2006) Identification of a common gene expression signature in dilated cardiomyopathy across independent microarray studies. Journal of the American College of Cardiology 48: 1610–1617. - PubMed
-
- Barth AS, Merk S, Arnoldi E, Zwermann L, Kloos P, et al. (2005) Reprogramming of the human atrial transcriptome in permanent atrial fibrillation: expression of a ventricular-like genomic signature. Circulation research 96: 1022–1029. - PubMed
-
- Hall JL, Grindle S, Han X, Fermin D, Park S, et al. (2004) Genomic profiling of the human heart before and after mechanical support with a ventricular assist device reveals alterations in vascular signaling networks. Physiological genomics 17: 283–291. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- U19 DK62434-08/DK/NIDDK NIH HHS/United States
- 5 T32 HD07165/HD/NICHD NIH HHS/United States
- T32 HD007165/HD/NICHD NIH HHS/United States
- R01 HD007857/HD/NICHD NIH HHS/United States
- U19 DK062434/DK/NIDDK NIH HHS/United States
- P30 HD024064/HD/NICHD NIH HHS/United States
- NHLBI R01 HL061483/PHS HHS/United States
- R01 HL061483/HL/NHLBI NIH HHS/United States
- HD08818-37/HD/NICHD NIH HHS/United States
- R01 HL089792/HL/NHLBI NIH HHS/United States
- CA125123/CA/NCI NIH HHS/United States
- P30 CA125123/CA/NCI NIH HHS/United States
- R01 HL022512/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

