Romulus and Remus, two phage isolates representing a distinct clade within the Twortlikevirus genus, display suitable properties for phage therapy applications
- PMID: 23302893
- PMCID: PMC3592175
- DOI: 10.1128/JVI.02763-12
Romulus and Remus, two phage isolates representing a distinct clade within the Twortlikevirus genus, display suitable properties for phage therapy applications
Abstract
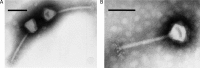
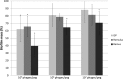
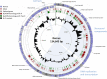
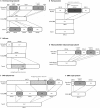
The renewed interest in controlling Staphylococcus aureus infections using their natural enemies, bacteriophages, has led to the isolation of a limited number of virulent phages so far. These phages are all members of the Twortlikevirus, displaying little variance. We present two novel closely related (95.9% DNA homology) lytic myoviruses, Romulus and Remus, with double-stranded DNA (dsDNA) genomes of 131,333 bp and 134,643 bp, respectively. Despite their relatedness to Staphylococcus phages K, G1, ISP, and Twort and Listeria phages A511 and P100, Romulus and Remus can be proposed as isolates of a new species within the Twortlikevirus genus. A distinguishing feature for these phage genomes is the unique distribution of group I introns compared to that in other staphylococcal myoviruses. In addition, a hedgehog/intein domain was found within their DNA polymerase genes, and an insertion sequence-encoded transposase exhibits splicing behavior and produces a functional portal protein. From a phage therapy application perspective, Romulus and Remus infected approximately 70% of the tested S. aureus isolates and displayed promising lytic activity against these isolates. Furthermore, both phages showed a rapid initial adsorption and demonstrated biofilm-degrading capacity in a proof-of-concept experiment.
Figures
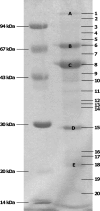
Similar articles
-
Comparative Genomics of Three Novel Jumbo Bacteriophages Infecting Staphylococcus aureus.J Virol. 2021 Sep 9;95(19):e0239120. doi: 10.1128/JVI.02391-20. Epub 2021 Sep 9. J Virol. 2021. PMID: 34287047 Free PMC article.
-
Characterization of vB_SauM-fRuSau02, a Twort-Like Bacteriophage Isolated from a Therapeutic Phage Cocktail.Viruses. 2017 Sep 14;9(9):258. doi: 10.3390/v9090258. Viruses. 2017. PMID: 28906479 Free PMC article.
-
Genomics of staphylococcal Twort-like phages--potential therapeutics of the post-antibiotic era.Adv Virus Res. 2012;83:143-216. doi: 10.1016/B978-0-12-394438-2.00005-0. Adv Virus Res. 2012. PMID: 22748811 Review.
-
Characterization and complete genome of the virulent Myoviridae phage JD007 active against a variety of Staphylococcus aureus isolates from different hospitals in Shanghai, China.Virol J. 2017 Feb 8;14(1):26. doi: 10.1186/s12985-017-0701-0. Virol J. 2017. PMID: 28179010 Free PMC article.
-
The SPO1-related bacteriophages.Arch Virol. 2010 Oct;155(10):1547-61. doi: 10.1007/s00705-010-0783-0. Epub 2010 Aug 17. Arch Virol. 2010. PMID: 20714761 Review.
Cited by
-
Characterization and Genome Analysis of Staphylococcus aureus Podovirus CSA13 and Its Anti-Biofilm Capacity.Viruses. 2019 Jan 12;11(1):54. doi: 10.3390/v11010054. Viruses. 2019. PMID: 30642091 Free PMC article.
-
Genomic analysis of Staphylococcus phage Stau2 isolated from medical specimen.Virus Genes. 2016 Feb;52(1):107-16. doi: 10.1007/s11262-015-1276-8. Epub 2015 Dec 26. Virus Genes. 2016. PMID: 26706853
-
Analysis of the virome associated to grapevine downy mildew lesions reveals new mycovirus lineages.Virus Evol. 2020 Nov 30;6(2):veaa058. doi: 10.1093/ve/veaa058. eCollection 2020 Jul. Virus Evol. 2020. PMID: 33324489 Free PMC article.
-
A blueprint for broadly effective bacteriophage-antibiotic cocktails against bacterial infections.Nat Commun. 2024 Nov 28;15(1):9987. doi: 10.1038/s41467-024-53994-9. Nat Commun. 2024. PMID: 39609398 Free PMC article.
-
An Appraisal of Bacteriophage Isolation Techniques from Environment.Microb Ecol. 2022 Apr;83(3):519-535. doi: 10.1007/s00248-021-01782-z. Epub 2021 Jun 17. Microb Ecol. 2022. PMID: 34136953 Review.
References
-
- Lowy FD. 1998. Staphylococcus aureus infections. N. Engl. J. Med. 339:520–532 - PubMed
-
- Veien NK. 1998. The clinician's choice of antibiotics in the treatment of bacterial skin infection. Br. J. Dermatol. 139:30–36 - PubMed
-
- Patel R. 2005. Biofilms and antimicrobial resistance. Clin. Orthop. Relat. Res. 437:41–47 - PubMed
-
- Klumpp J, Lavigne R, Loessner MJ, Ackermann HW. 2010. The SPO1-related bacteriophages. Arch. Virol. 155:1547–1561 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

