Gene-specific factors determine mitotic expression and bookmarking via alternate regulatory elements
- PMID: 23303784
- PMCID: PMC4230186
- DOI: 10.1093/nar/gks1365
Gene-specific factors determine mitotic expression and bookmarking via alternate regulatory elements
Abstract
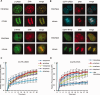
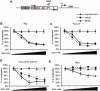
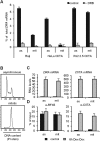
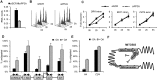
Transcriptional silencing during mitosis is caused by inactivation of critical transcriptional regulators and/or chromatin condensation. Inheritance of gene expression patterns through cell division involves various bookmarking mechanisms. In this report, we have examined the mitotic and post-mitotic expression of the DRA major histocompatibility class II (MHCII) gene in different cell types. During mitosis the constitutively MHCII-expressing B lymphoblastoid cells showed sustained occupancy of the proximal promoter by the cognate enhanceosome and general transcription factors. In contrast, although mitotic epithelial cells were depleted of these proteins irrespectively of their MHCII transcriptional activity, a distal enhancer selectively recruited the PP2A phosphatase via NFY and maintained chromatin accessibility. Based on our data, we propose a novel chromatin anti-condensation role for this element in mitotic bookmarking and timing of post-mitotic transcriptional reactivation.
Figures



Similar articles
-
A hyperactive transcriptional state marks genome reactivation at the mitosis-G1 transition.Genes Dev. 2016 Jun 15;30(12):1423-39. doi: 10.1101/gad.280859.116. Genes Dev. 2016. PMID: 27340175 Free PMC article.
-
Identification of GA-Binding Protein Transcription Factor Alpha Subunit (GABPA) as a Novel Bookmarking Factor.Int J Mol Sci. 2019 Mar 4;20(5):1093. doi: 10.3390/ijms20051093. Int J Mol Sci. 2019. PMID: 30836589 Free PMC article.
-
Kaposi's Sarcoma-Associated Herpesvirus Latency-Associated Nuclear Antigen Inhibits Major Histocompatibility Complex Class II Expression by Disrupting Enhanceosome Assembly through Binding with the Regulatory Factor X Complex.J Virol. 2015 May;89(10):5536-56. doi: 10.1128/JVI.03713-14. Epub 2015 Mar 4. J Virol. 2015. PMID: 25740990 Free PMC article.
-
Mitotic bookmarking in development and stem cells.Development. 2017 Oct 15;144(20):3633-3645. doi: 10.1242/dev.146522. Development. 2017. PMID: 29042475 Review.
-
A changing paradigm of transcriptional memory propagation through mitosis.Nat Rev Mol Cell Biol. 2019 Jan;20(1):55-64. doi: 10.1038/s41580-018-0077-z. Nat Rev Mol Cell Biol. 2019. PMID: 30420736 Free PMC article. Review.
Cited by
-
Nuclear organization mediates cancer-compromised genetic and epigenetic control.Adv Biol Regul. 2018 Aug;69:1-10. doi: 10.1016/j.jbior.2018.05.001. Epub 2018 May 9. Adv Biol Regul. 2018. PMID: 29759441 Free PMC article. Review.
-
A high definition look at the NF-Y regulome reveals genome-wide associations with selected transcription factors.Nucleic Acids Res. 2016 Jun 2;44(10):4684-702. doi: 10.1093/nar/gkw096. Epub 2016 Feb 20. Nucleic Acids Res. 2016. PMID: 26896797 Free PMC article.
-
Inhibition of the RUNX1-CBFβ transcription factor complex compromises mammary epithelial cell identity: a phenotype potentially stabilized by mitotic gene bookmarking.Oncotarget. 2020 Jun 30;11(26):2512-2530. doi: 10.18632/oncotarget.27637. eCollection 2020 Jun 30. Oncotarget. 2020. PMID: 32655837 Free PMC article.
-
Promyelocytic leukemia protein (PML) controls breast cancer cell proliferation by modulating Forkhead transcription factors.Mol Oncol. 2019 Jun;13(6):1369-1387. doi: 10.1002/1878-0261.12486. Epub 2019 May 16. Mol Oncol. 2019. PMID: 30927552 Free PMC article.
-
Acute Stress Enhances Epigenetic Modifications But Does Not Affect the Constitutive Binding of pCREB to Immediate-Early Gene Promoters in the Rat Hippocampus.Front Mol Neurosci. 2017 Dec 19;10:416. doi: 10.3389/fnmol.2017.00416. eCollection 2017. Front Mol Neurosci. 2017. PMID: 29311809 Free PMC article.
References
-
- Reith W, Mach B. The bare lymphocyte syndrome and the regulation of MHC expression. Annu. Rev. Immunol. 2001;19:331–373. - PubMed
-
- Nagarajan UM, Louis-Plence P, DeSandro A, Nilsen R, Bushey A, Boss JM. RFX-B is the gene responsible for the most common cause of the bare lymphocyte syndrome, an MHC class II immunodeficiency. Immunity. 1999;10:153–162. - PubMed
-
- Steimle V, Durand B, Barras E, Zufferey M, Hadam MR, Mach B, Reith W. A novel DNA-binding regulatory factor is mutated in primary MHC class II deficiency (bare lymphocyte syndrome) Genes Dev. 1995;9:1021–1032. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

