RIM3γ and RIM4γ are key regulators of neuronal arborization
- PMID: 23303958
- PMCID: PMC6704911
- DOI: 10.1523/JNEUROSCI.2229-12.2013
RIM3γ and RIM4γ are key regulators of neuronal arborization
Abstract
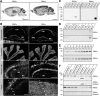
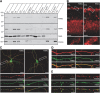
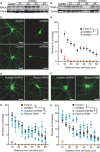
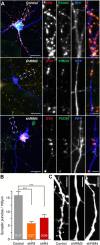
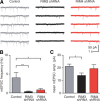
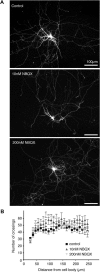
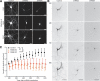
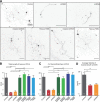
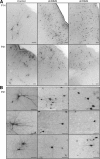
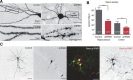
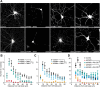
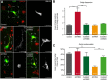
The large isoforms of the Rab3 interacting molecule (RIM) family, RIM1α/β and RIM2α/β, have been shown to be centrally involved in mediating presynaptic active zone function. The RIM protein family contains two additional small isoforms, RIM3γ and RIM4γ, which are composed only of the RIM-specific C-terminal C2B domain and varying N-terminal sequences and whose function remains to be elucidated. Here, we report that both, RIM3γ and RIM4γ, play an essential role for the development of neuronal arborization and of dendritic spines independent of synaptic function. γ-RIM knock-down in rat primary neuronal cultures and in vivo resulted in a drastic reduction in the complexity of neuronal arborization, affecting both axonal and dendritic outgrowth, independent of the time point of γ-RIM downregulation during dendrite development. Rescue experiments revealed that the phenotype is caused by a function common to both γ-RIMs. These findings indicate that γ-RIMs are involved in cell biological functions distinct from the regulation of synaptic vesicle exocytosis and play a role in the molecular mechanisms controlling the establishment of dendritic complexity and axonal outgrowth.
Figures
References
-
- Betz A, Thakur P, Junge HJ, Ashery U, Rhee JS, Scheuss V, Rosenmund C, Rettig J, Brose N. Functional interaction of the active zone proteins Munc13-1 and RIM1 in synaptic vesicle priming. Neuron. 2001;30:183–196. - PubMed
-
- Calakos N, Schoch S, Südhof TC, Malenka RC. Multiple roles for the active zone protein RIM1alpha in late stages of neurotransmitter release. Neuron. 2004;42:889–896. - PubMed
-
- Coppola T, Magnin-Luthi S, Perret-Menoud V, Gattesco S, Schiavo G, Regazzi R. Direct interaction of the Rab3 effector RIM with Ca2+ channels, SNAP-25, and synaptotagmin. J Biol Chem. 2001;276:32756–32762. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
