Fitness components and natural selection: why are there different patterns on the emergence of drug resistance in Plasmodium falciparum and Plasmodium vivax?
- PMID: 23305428
- PMCID: PMC3571882
- DOI: 10.1186/1475-2875-12-15
Fitness components and natural selection: why are there different patterns on the emergence of drug resistance in Plasmodium falciparum and Plasmodium vivax?
Abstract
Background: Considering the distinct biological characteristics of Plasmodium species is crucial for control and elimination efforts, in particular when facing the spread of drug resistance. Whereas the evolutionary fitness of all malarial species could be approximated by the probability of being taken by a mosquito and then infecting a new host, the actual steps in the malaria life cycle leading to a successful transmission event show differences among Plasmodium species. These "steps" are called fitness components. Differences in terms of fitness components may affect how selection imposed by interventions, e.g. drug treatments, differentially acts on each Plasmodium species. Thus, a successful malaria control or elimination programme should understand how differences in fitness components among different malaria species could affect adaptive evolution (e.g. the emergence of drug resistance). In this investigation, the interactions between some fitness components and natural selection are explored.
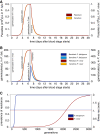
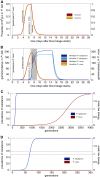
Methods: A population-genetic model is formulated that qualitatively explains how different fitness components (in particular gametocytogenesis and longevity of gametocytes) affect selection acting on merozoites during the erythrocytic cycle. By comparing Plasmodium falciparum and Plasmodium vivax, the interplay of parasitaemia and gametocytaemia dynamics in determining fitness is modelled under circumstances that allow contrasting solely the differences between these two parasites in terms of their fitness components.
Results: By simulating fitness components, it is shown that selection acting on merozoites (e.g., on drug resistant mutations or malaria antigens) is more efficient in P. falciparum than in P. vivax. These results could explain, at least in part, why resistance against drugs, such as chloroquine (CQ) is highly prevalent in P. falciparum worldwide, while CQ is still a successful treatment for P. vivax despite its massive use. Furthermore, these analyses are used to explore the importance of understanding the dynamic of gametocytaemia to ascertain the spreading of drug resistance.
Conclusions: The strength of natural selection on mutations that express their advantage at the merozoite stage is different in P. vivax and P. falciparum. Species-specific differences in gametocytogenesis and longevity of gametocytes need to be accounted for when designing effective malaria control and elimination programmes. There is a need for reliable data on gametocytogenesis from field studies.
Figures
Similar articles
-
Detection of high levels of mutations involved in anti-malarial drug resistance in Plasmodium falciparum and Plasmodium vivax at a rural hospital in southern Ethiopia.Malar J. 2011 Aug 2;10:214. doi: 10.1186/1475-2875-10-214. Malar J. 2011. PMID: 21810256 Free PMC article.
-
Prevalence of molecular markers of anti-malarial drug resistance in Plasmodium vivax and Plasmodium falciparum in two districts of Nepal.Malar J. 2011 Apr 1;10:75. doi: 10.1186/1475-2875-10-75. Malar J. 2011. PMID: 21457533 Free PMC article.
-
Susceptibility of Plasmodium falciparum to artemisinins and Plasmodium vivax to chloroquine in Phuoc Chien Commune, Ninh Thuan Province, south-central Vietnam.Malar J. 2019 Jan 17;18(1):10. doi: 10.1186/s12936-019-2640-2. Malar J. 2019. PMID: 30654808 Free PMC article.
-
Epidemiology and infectivity of Plasmodium falciparum and Plasmodium vivax gametocytes in relation to malaria control and elimination.Clin Microbiol Rev. 2011 Apr;24(2):377-410. doi: 10.1128/CMR.00051-10. Clin Microbiol Rev. 2011. PMID: 21482730 Free PMC article. Review.
-
[Mechanisms and dynamics of drug resistance in Plasmodium falciparum].Bull Soc Pathol Exot. 1999 Sep-Oct;92(4):236-41. Bull Soc Pathol Exot. 1999. PMID: 10572658 Review. French.
Cited by
-
The evolution and diversity of a low complexity vaccine candidate, merozoite surface protein 9 (MSP-9), in Plasmodium vivax and closely related species.Infect Genet Evol. 2013 Dec;20:239-48. doi: 10.1016/j.meegid.2013.09.011. Epub 2013 Sep 14. Infect Genet Evol. 2013. PMID: 24044894 Free PMC article.
-
Whole-genome sequencing of a Plasmodium vivax clinical isolate exhibits geographical characteristics and high genetic variation in China-Myanmar border area.BMC Genomics. 2017 Feb 6;18(1):131. doi: 10.1186/s12864-017-3523-y. BMC Genomics. 2017. PMID: 28166727 Free PMC article.
-
Genome-wide patterns of genetic polymorphism and signatures of selection in Plasmodium vivax.Genome Biol Evol. 2014 Dec 17;7(1):106-19. doi: 10.1093/gbe/evu267. Genome Biol Evol. 2014. PMID: 25523904 Free PMC article.
-
Investigation of Mutations in the crt-o and mdr1 Genes of Plasmodium vivax for the Molecular Surveillance of Chloroquine Resistance in Parasites from Gold Mining Areas in Roraima, Brazil.Microorganisms. 2024 Aug 15;12(8):1680. doi: 10.3390/microorganisms12081680. Microorganisms. 2024. PMID: 39203521 Free PMC article.
-
Genomic Analysis Reveals a Common Breakpoint in Amplifications of the Plasmodium vivax Multidrug Resistance 1 Locus in Thailand.J Infect Dis. 2016 Oct 15;214(8):1235-42. doi: 10.1093/infdis/jiw323. Epub 2016 Jul 24. J Infect Dis. 2016. PMID: 27456706 Free PMC article.
References
-
- Coatney G, Collins W, Warren M, Contacos P. The primate malarias. Bethesda: U.S. National Institute of Allergy and Infectious Diseases; 1971.
-
- Liu W, Li Y, Learn GH, Rudicell RS, Robertson JD, Keele BF, Ndjango JB, Sanz CM, Morgan DB, Locatelli S, Gonder MK, Kranzusch PJ, Walsh PD, Delaporte E, Mpoudi-Ngole E, Georgiev AV, Muller MN, Shaw GM, Peeters M, Sharp PM, Rayner JC, Hahn BH. Origin of the human malaria parasite Plasmodium falciparum in gorillas. Nature. 2010;467:420–425. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

