Fine mapping links the FTa1 flowering time regulator to the dominant spring1 locus in Medicago
- PMID: 23308229
- PMCID: PMC3538541
- DOI: 10.1371/journal.pone.0053467
Fine mapping links the FTa1 flowering time regulator to the dominant spring1 locus in Medicago
Abstract
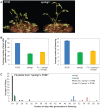
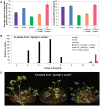
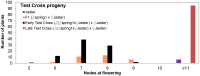
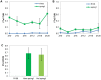
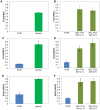
To extend our understanding of flowering time control in eudicots, we screened for mutants in the model legume Medicago truncatula (Medicago). We identified an early flowering mutant, spring1, in a T-DNA mutant screen, but spring1 was not tagged and was deemed a somaclonal mutant. We backcrossed the mutant to wild type R108. The F1 plants and the majority of F2 plants were early flowering like spring1, strongly indicating that spring1 conferred monogenic, dominant early flowering. We hypothesized that the spring1 phenotype resulted from over expression of an activator of flowering. Previously, a major QTL for flowering time in different Medicago accessions was located to an interval on chromosome 7 with six candidate flowering-time activators, including a CONSTANS gene, MtCO, and three FLOWERING LOCUS T (FT) genes. Hence we embarked upon linkage mapping using 29 markers from the MtCO/FT region on chromosome 7 on two populations developed by crossing spring1 with Jester. Spring1 mapped to an interval of ∼0.5 Mb on chromosome 7 that excluded MtCO, but contained 78 genes, including the three FT genes. Of these FT genes, only FTa1 was up-regulated in spring1 plants. We then investigated global gene expression in spring1 and R108 by microarray analysis. Overall, they had highly similar gene expression and apart from FTa1, no genes in the mapping interval were differentially expressed. Two MADS transcription factor genes, FRUITFULLb (FULb) and SUPPRESSOR OF OVER EXPRESSION OF CONSTANS1a (SOC1a), that were up-regulated in spring1, were also up-regulated in transgenic Medicago over-expressing FTa1. This suggested that their differential expression in spring1 resulted from the increased abundance of FTa1. A 6255 bp genomic FTa1 fragment, including the complete 5' region, was sequenced, but no changes were observed indicating that the spring1 mutation is not a DNA sequence difference in the FTa1 promoter or introns.
Conflict of interest statement
Figures

Similar articles
-
Medicago PHYA promotes flowering, primary stem elongation and expression of flowering time genes in long days.BMC Plant Biol. 2020 Jul 11;20(1):329. doi: 10.1186/s12870-020-02540-y. BMC Plant Biol. 2020. PMID: 32652925 Free PMC article.
-
Retroelement insertions at the Medicago FTa1 locus in spring mutants eliminate vernalisation but not long-day requirements for early flowering.Plant J. 2013 Nov;76(4):580-91. doi: 10.1111/tpj.12315. Epub 2013 Oct 5. Plant J. 2013. PMID: 23964816
-
MtVRN2 is a Polycomb VRN2-like gene which represses the transition to flowering in the model legume Medicago truncatula.Plant J. 2016 Apr;86(2):145-60. doi: 10.1111/tpj.13156. Plant J. 2016. PMID: 26947149
-
The Medicago FLOWERING LOCUS T homolog, MtFTa1, is a key regulator of flowering time.Plant Physiol. 2011 Aug;156(4):2207-24. doi: 10.1104/pp.111.180182. Epub 2011 Jun 17. Plant Physiol. 2011. PMID: 21685176 Free PMC article.
-
Functional Genomics and Flowering Time in Medicago truncatula: An Overview.Methods Mol Biol. 2018;1822:261-271. doi: 10.1007/978-1-4939-8633-0_17. Methods Mol Biol. 2018. PMID: 30043309 Review.
Cited by
-
A SOC1-like gene MtSOC1a promotes flowering and primary stem elongation in Medicago.J Exp Bot. 2018 Sep 14;69(20):4867-4880. doi: 10.1093/jxb/ery284. J Exp Bot. 2018. PMID: 30295903 Free PMC article.
-
Isolation and Functional Characterization of MsFTa, a FLOWERING LOCUS T Homolog from Alfalfa (Medicago sativa).Int J Mol Sci. 2019 Apr 22;20(8):1968. doi: 10.3390/ijms20081968. Int J Mol Sci. 2019. PMID: 31013631 Free PMC article.
-
Gene-edited Mtsoc1 triple mutant Medicago plants do not flower.Front Plant Sci. 2024 Feb 26;15:1357924. doi: 10.3389/fpls.2024.1357924. eCollection 2024. Front Plant Sci. 2024. PMID: 38469328 Free PMC article.
-
Genetic analysis of global faba bean diversity, agronomic traits and selection signatures.Theor Appl Genet. 2023 Apr 19;136(5):114. doi: 10.1007/s00122-023-04360-8. Theor Appl Genet. 2023. PMID: 37074596 Free PMC article.
-
Medicago PHYA promotes flowering, primary stem elongation and expression of flowering time genes in long days.BMC Plant Biol. 2020 Jul 11;20(1):329. doi: 10.1186/s12870-020-02540-y. BMC Plant Biol. 2020. PMID: 32652925 Free PMC article.
References
-
- Putterill J, Laurie R, Macknight R (2004) It´s time to flower: the genetic control of flowering time. BioEssays 26: 363–373. - PubMed
-
- Turck F, Fornara F, Coupland G (2008) Regulation and identity of florigen: FLOWERING LOCUS T moves center stage. Annual Review of Plant Biology 59: 573–594. - PubMed
-
- Turnbull C (2011) Long-distance regulation of flowering time. Journal of Experimental Botany 62: 4399–4413. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

