LPA1-induced cytoskeleton reorganization drives fibrosis through CTGF-dependent fibroblast proliferation
- PMID: 23322166
- PMCID: PMC3633809
- DOI: 10.1096/fj.12-219378
LPA1-induced cytoskeleton reorganization drives fibrosis through CTGF-dependent fibroblast proliferation
Abstract
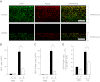
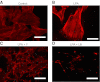
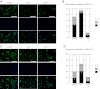
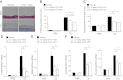
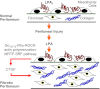
There has been much recent interest in lysophosphatidic acid (LPA) signaling through one of its receptors, LPA1, in fibrotic diseases, but the mechanisms by which LPA-LPA1 signaling promotes pathological fibrosis remain to be fully elucidated. Using a mouse peritoneal fibrosis model, we demonstrate central roles for LPA and LPA1 in fibroblast proliferation. Genetic deletion or pharmacological antagonism of LPA1 protected mice from peritoneal fibrosis, blunting the increases in peritoneal collagen by 65.4 and 52.9%, respectively, compared to control animals and demonstrated that peritoneal fibroblast proliferation was highly LPA1 dependent. Activation of LPA1 on mesothelial cells induced these cells to express connective tissue growth factor (CTGF), driving fibroblast proliferation in a paracrine fashion. Activation of mesothelial cell LPA1 induced CTGF expression by inducing cytoskeleton reorganization in these cells, causing nuclear translocation of myocardin-related transcription factor (MRTF)-A and MRTF-B. Pharmacological inhibition of MRTF-induced transcription also diminished CTGF expression and fibrosis in the peritoneal fibrosis model, mitigating the increase in peritoneal collagen content by 57.9% compared to controls. LPA1-induced cytoskeleton reorganization therefore makes a previously unrecognized but critically important contribution to the profibrotic activities of LPA by driving MRTF-dependent CTGF expression, which, in turn, drives fibroblast proliferation.
Figures




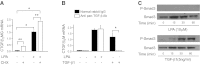


References
-
- Selman M., King T. E., Pardo A. (2001) Idiopathic pulmonary fibrosis: prevailing and evolving hypotheses about its pathogenesis and implications for therapy. Ann. Intern. Med. 134, 136–151 - PubMed
-
- Scotton C. J., Chambers R. C. (2007) Molecular targets in pulmonary fibrosis: the myofibroblast in focus. Chest 132, 1311–1321 - PubMed
-
- Choi J. W., Herr D. R., Noguchi K., Yung Y. C., Lee C. W., Mutoh T., Lin M. E., Teo S. T., Park K. E., Mosley A. N., Chun J. (2010) LPA receptors: subtypes and biological actions. Annu. Rev. Pharmacol. Toxicol. 50, 157–186 - PubMed
-
- Tager A. M., LaCamera P., Shea B. S., Campanella G. S., Selman M., Zhao Z., Polosukhin V., Wain J., Karimi-Shah B. A., Kim N. D., Hart W. K., Pardo A., Blackwell T. S., Xu Y., Chun J., Luster A. D. (2008) The lysophosphatidic acid receptor LPA1 links pulmonary fibrosis to lung injury by mediating fibroblast recruitment and vascular leak. Nat. Med. 14, 45–54 - PubMed
-
- Castelino F. V., Seiders J., Bain G., Brooks S. F., King C. D., Swaney J. S., Lorrain D. S., Chun J., Luster A. D., Tager A. M. (2011) Amelioration of dermal fibrosis by genetic deletion or pharmacologic antagonism of lysophosphatidic acid receptor 1 in a mouse model of scleroderma. Arthritis Rheum. 63, 1405–1415 - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

