Architecture and assembly of HIV integrase multimers in the absence of DNA substrates
- PMID: 23322775
- PMCID: PMC3591645
- DOI: 10.1074/jbc.M112.434431
Architecture and assembly of HIV integrase multimers in the absence of DNA substrates
Abstract
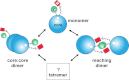
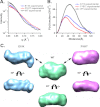
We have applied small angle x-ray scattering and protein cross-linking coupled with mass spectrometry to determine the architectures of full-length HIV integrase (IN) dimers in solution. By blocking interactions that stabilize either a core-core domain interface or N-terminal domain intermolecular contacts, we show that full-length HIV IN can form two dimer types. One is an expected dimer, characterized by interactions between two catalytic core domains. The other dimer is stabilized by interactions of the N-terminal domain of one monomer with the C-terminal domain and catalytic core domain of the second monomer as well as direct interactions between the two C-terminal domains. This organization is similar to the "reaching dimer" previously described for wild type ASV apoIN and resembles the inner, substrate binding dimer in the crystal structure of the PFV intasome. Results from our small angle x-ray scattering and modeling studies indicate that in the absence of its DNA substrate, the HIV IN tetramer assembles as two stacked reaching dimers that are stabilized by core-core interactions. These models of full-length HIV IN provide new insight into multimer assembly and suggest additional approaches for enzyme inhibition.
Figures



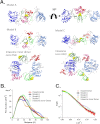


References
-
- Bojja R. S., Andrake M. D., Weigand S., Merkel G., Yarychkivska O., Henderson A., Kummerling M., Skalka A. M. (2011) Architecture of a full-length retroviral integrase monomer and dimer, revealed by small angle x-ray scattering and chemical cross-linking. J. Biol. Chem. 286, 17047–17059 - PMC - PubMed
-
- Grabarek Z., Gergely J. (1990) Zero-length cross-linking procedure with the use of active esters. Anal. Biochem. 185, 131–135 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

