The complex transcriptional landscape of the anucleate human platelet
- PMID: 23323973
- PMCID: PMC3722126
- DOI: 10.1186/1471-2164-14-1
The complex transcriptional landscape of the anucleate human platelet
Abstract
Background: Human blood platelets are essential to maintaining normal hemostasis, and platelet dysfunction often causes bleeding or thrombosis. Estimates of genome-wide platelet RNA expression using microarrays have provided insights to the platelet transcriptome but were limited by the number of known transcripts. The goal of this effort was to deep-sequence RNA from leukocyte-depleted platelets to capture the complex profile of all expressed transcripts.
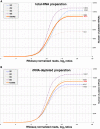
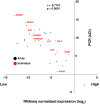
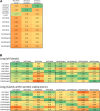
Results: From each of four healthy individuals we generated long RNA (≥40 nucleotides) profiles from total and ribosomal-RNA depleted RNA preparations, as well as short RNA (<40 nucleotides) profiles. Analysis of ~1 billion reads revealed that coding and non-coding platelet transcripts span a very wide dynamic range (≥16 PCR cycles beyond β-actin), a result we validated through qRT-PCR on many dozens of platelet messenger RNAs. Surprisingly, ribosomal-RNA depletion significantly and adversely affected estimates of the relative abundance of transcripts. Of the known protein-coding loci, ~9,500 are present in human platelets. We observed a strong correlation between mRNAs identified by RNA-seq and microarray for well-expressed mRNAs, but RNASeq identified many more transcripts of lower abundance and permitted discovery of novel transcripts.
Conclusions: Our analyses revealed diverse classes of non-coding RNAs, including: pervasive antisense transcripts to protein-coding loci; numerous, previously unreported and abundant microRNAs; retrotransposons; and thousands of novel un-annotated long and short intronic transcripts, an intriguing finding considering the anucleate nature of platelets. The data are available through a local mirror of the UCSC genome browser and can be accessed at: http://cm.jefferson.edu/platelets_2012/.
Figures

References
-
- Jones CI, Garner SF, Angenent W, Bernard A, Berzuini C, Burns P, Farndale RW, Hogwood J, Rankin A, Stephens JC. et al. Mapping the platelet profile for functional genomic studies and demonstration of the effect size of the GP6 locus. Journal of thrombosis and haemostasis: JTH. 2007;5(8):1756–1765. doi: 10.1111/j.1538-7836.2007.02632.x. - DOI - PubMed
-
- Bray PF, Mathias RA, Faraday N, Yanek LR, Fallin MD, Herrera-Galeano JE, Wilson AF, Becker LC, Becker DM. Heritability of platelet function in families with premature coronary artery disease. Journal of thrombosis and haemostasis: JTH. 2007;5(8):1617–1623. doi: 10.1111/j.1538-7836.2007.02618.x. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

