A RAD-based linkage map and comparative genomics in the gudgeons (genus Gnathopogon, Cyprinidae)
- PMID: 23324215
- PMCID: PMC3583795
- DOI: 10.1186/1471-2164-14-32
A RAD-based linkage map and comparative genomics in the gudgeons (genus Gnathopogon, Cyprinidae)
Abstract
Background: The construction of linkage maps is a first step in exploring the genetic basis for adaptive phenotypic divergence in closely related species by quantitative trait locus (QTL) analysis. Linkage maps are also useful for comparative genomics in non-model organisms. Advances in genomics technologies make it more feasible than ever to study the genetics of adaptation in natural populations. Restriction-site associated DNA (RAD) sequencing in next-generation sequencers facilitates the development of many genetic markers and genotyping. We aimed to construct a linkage map of the gudgeons of the genus Gnathopogon (Cyprinidae) for comparative genomics with the zebrafish Danio rerio (a member of the same family as gudgeons) and for the future QTL analysis of the genetic architecture underlying adaptive phenotypic evolution of Gnathopogon.
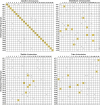
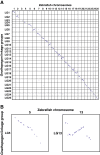
Results: We constructed the first genetic linkage map of Gnathopogon using a 198 F2 interspecific cross between two closely related species in Japan: river-dwelling Gnathopogon elongatus and lake-dwelling Gnathopogon caerulescens. Based on 1,622 RAD-tag markers, a linkage map spanning 1,390.9 cM with 25 linkage groups and an average marker interval of 0.87 cM was constructed. We also identified a region involving female-specific transmission ratio distortion (TRD). Synteny and collinearity were extensively conserved between Gnathopogon and zebrafish.
Conclusions: The dense SNP-based linkage map presented here provides a basis for future QTL analysis. It will also be useful for transferring genomic information from a "traditional" model fish species, zebrafish, to screen candidate genes underlying ecologically important traits of the gudgeons.
Figures
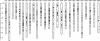


References
-
- Schluter D. The Ecology of Adaptive Radiation. Oxford: Oxford University Press; 2000.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

