Screen for Footprints of Selection during Domestication/Captive Breeding of Atlantic Salmon
- PMID: 23326209
- PMCID: PMC3544263
- DOI: 10.1155/2012/628204
Screen for Footprints of Selection during Domestication/Captive Breeding of Atlantic Salmon
Abstract
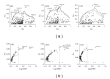
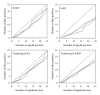
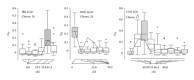
Domesticated animals provide a unique opportunity to identify genomic targets of artificial selection to the captive environment. Here, we screened three independent domesticated/captive Atlantic salmon (Salmo salar) strains and their wild progenitor populations in an effort to detect potential signals of domestication selection by typing of 261 SNPs and 70 microsatellite loci. By combining information from four different neutrality tests, in total ten genomic regions showed signs of directional selection based on multiple sources of evidence. Most of the identified candidate regions were rather small ranging from zero to a few centimorgans (cM) in the female Atlantic salmon linkage map. We also evaluated how adaptation from standing variation affects adjacent SNP and microsatellite variation along the chromosomes and, by using forward simulations with strong selection, we were able to generate genetic differentiation patterns comparable to the observed data. This study highlights the significance of standing genetic variation during the early stages of adaptation and represents a useful step towards identifying functional variants involved in domestication of Atlantic salmon.
Figures


Similar articles
-
Comparing genomic signatures of domestication in two Atlantic salmon (Salmo salar L.) populations with different geographical origins.Evol Appl. 2018 Dec 7;12(1):137-156. doi: 10.1111/eva.12689. eCollection 2019 Jan. Evol Appl. 2018. PMID: 30622641 Free PMC article.
-
Population genomic analyses of early-phase Atlantic Salmon (Salmo salar) domestication/captive breeding.Evol Appl. 2015 Jan;8(1):93-107. doi: 10.1111/eva.12230. Epub 2014 Nov 20. Evol Appl. 2015. PMID: 25667605 Free PMC article.
-
Parallel Selection in Domesticated Atlantic Salmon from Divergent Founders Including on Whole-Genome Duplication-derived Homeologous Regions.Genome Biol Evol. 2025 Apr 3;17(4):evaf063. doi: 10.1093/gbe/evaf063. Genome Biol Evol. 2025. PMID: 40247730 Free PMC article.
-
Footprints of directional selection in wild Atlantic salmon populations: evidence for parasite-driven evolution?PLoS One. 2014 Mar 26;9(3):e91672. doi: 10.1371/journal.pone.0091672. eCollection 2014. PLoS One. 2014. PMID: 24670947 Free PMC article.
-
A critical review of adaptive genetic variation in Atlantic salmon: implications for conservation.Biol Rev Camb Philos Soc. 2007 May;82(2):173-211. doi: 10.1111/j.1469-185X.2006.00004.x. Biol Rev Camb Philos Soc. 2007. PMID: 17437557 Review.
Cited by
-
Changed Patterns of Genomic Variation Following Recent Domestication: Selection Sweeps in Farmed Atlantic Salmon.Front Genet. 2020 Apr 3;11:264. doi: 10.3389/fgene.2020.00264. eCollection 2020. Front Genet. 2020. PMID: 32318091 Free PMC article.
-
Consequences of domestication in eastern oyster: Insights from whole genomic analyses.Evol Appl. 2024 May 29;17(6):e13710. doi: 10.1111/eva.13710. eCollection 2024 Jun. Evol Appl. 2024. PMID: 38817396 Free PMC article.
-
A genome scan for selection signatures comparing farmed Atlantic salmon with two wild populations: Testing colocalization among outlier markers, candidate genes, and quantitative trait loci for production traits.Evol Appl. 2016 Dec 29;10(3):276-296. doi: 10.1111/eva.12450. eCollection 2017 Mar. Evol Appl. 2016. PMID: 28250812 Free PMC article.
-
Comparing genomic signatures of domestication in two Atlantic salmon (Salmo salar L.) populations with different geographical origins.Evol Appl. 2018 Dec 7;12(1):137-156. doi: 10.1111/eva.12689. eCollection 2019 Jan. Evol Appl. 2018. PMID: 30622641 Free PMC article.
-
Genetic Variation and Breeding Signature in Mass Selection Lines of the Pacific Oyster (Crassostrea gigas) Assessed by SNP Markers.PLoS One. 2016 Mar 8;11(3):e0150868. doi: 10.1371/journal.pone.0150868. eCollection 2016. PLoS One. 2016. PMID: 26954577 Free PMC article.
References
-
- Purugganan MD, Fuller DQ. The nature of selection during plant domestication. Nature. 2009;457(7231):843–848. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

