Gene expression-based classifiers identify Staphylococcus aureus infection in mice and humans
- PMID: 23326304
- PMCID: PMC3541361
- DOI: 10.1371/journal.pone.0048979
Gene expression-based classifiers identify Staphylococcus aureus infection in mice and humans
Abstract
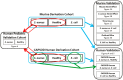
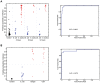
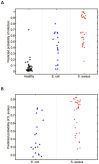
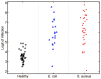
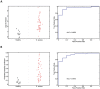
Staphylococcus aureus causes a spectrum of human infection. Diagnostic delays and uncertainty lead to treatment delays and inappropriate antibiotic use. A growing literature suggests the host's inflammatory response to the pathogen represents a potential tool to improve upon current diagnostics. The hypothesis of this study is that the host responds differently to S. aureus than to E. coli infection in a quantifiable way, providing a new diagnostic avenue. This study uses Bayesian sparse factor modeling and penalized binary regression to define peripheral blood gene-expression classifiers of murine and human S. aureus infection. The murine-derived classifier distinguished S. aureus infection from healthy controls and Escherichia coli-infected mice across a range of conditions (mouse and bacterial strain, time post infection) and was validated in outbred mice (AUC>0.97). A S. aureus classifier derived from a cohort of 94 human subjects distinguished S. aureus blood stream infection (BSI) from healthy subjects (AUC 0.99) and E. coli BSI (AUC 0.84). Murine and human responses to S. aureus infection share common biological pathways, allowing the murine model to classify S. aureus BSI in humans (AUC 0.84). Both murine and human S. aureus classifiers were validated in an independent human cohort (AUC 0.95 and 0.92, respectively). The approach described here lends insight into the conserved and disparate pathways utilized by mice and humans in response to these infections. Furthermore, this study advances our understanding of S. aureus infection; the host response to it; and identifies new diagnostic and therapeutic avenues.
Conflict of interest statement
Figures

References
-
- Martin GS, Mannino DM, Eaton S, Moss M (2003) The epidemiology of sepsis in the United States from 1979 through 2000. N Engl J Med 348: 1546–1554. - PubMed
-
- Kollef MH, Sherman G, Ward S, Fraser VJ (1999) Inadequate antimicrobial treatment of infections: a risk factor for hospital mortality among critically ill patients. Chest 115: 462–474. - PubMed
-
- Kumar A, Ellis P, Arabi Y, Roberts D, Light B, et al. (2009) Initiation of inappropriate antimicrobial therapy results in a fivefold reduction of survival in human septic shock. Chest 136: 1237–1248. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

