The transcription factor Sp3 regulates the expression of a metastasis-related marker of sarcoma, actin filament-associated protein 1-like 1 (AFAP1L1)
- PMID: 23326307
- PMCID: PMC3541374
- DOI: 10.1371/journal.pone.0049709
The transcription factor Sp3 regulates the expression of a metastasis-related marker of sarcoma, actin filament-associated protein 1-like 1 (AFAP1L1)
Abstract
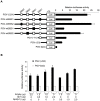
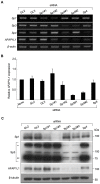
We previously identified actin filament-associated protein 1-like 1 (AFAP1L1) as a metastasis-predicting marker from the gene-expression profiles of 65 spindle cell sarcomas, and demonstrated the up-regulation of AFAP1L1 expression to be an independent risk factor for distant metastasis in multivariate analyses. Little is known, however, about how the expression of AFAP1L1 is regulated. Luciferase reporter assays showed tandem binding motives of a specificity protein (Sp) located at -85 to -75 relative to the transcriptional start site to be essential to the promoter activity. Overexpression of Sp1 and Sp3 proteins transactivated the proximal AFAP1L1 promoter construct, and electrophoretic mobility shift assays showed that both Sp1 and Sp3 were able to bind to this region in vitro. Chromatin immunoprecipitation experiments, however, revealed that Sp3 is the major factor binding to the proximal promoter region of the AFAP1L1 gene in AFAP1L1- positive cells. Treatment with mithramycin A, an inhibitor of proteins binding to GC-rich regions, prevented Sp3 from binding to the proximal promoter region of AFAP1L1 and decreased its expression in a dose-dependent manner. Finally, knocking down Sp3 using small inhibitory RNA duplex (siRNA) reduced AFAP1L1 expression significantly, which was partially restored by expressing siRNA-resistant Sp3. These findings indicate a novel role for Sp3 in sarcomas as a driver for expression of the metastasis-related gene AFAP1L1.
Conflict of interest statement
Figures






References
-
- Furu M, Kajita Y, Nagayama S, Ishibe T, Shima Y, et al. (2011) Identification of AFAP1L1 as a prognostic marker for spindle cell sarcomas. Oncogene 30: 4015–4025. - PubMed
-
- Kawai A, Naito N, Yoshida A, Morimoto Y, Ouchida M, et al. (2004) Establishment and characterization of a biphasic synovial sarcoma cell line, SYO-1. Cancer Lett 204: 105–113. - PubMed
-
- Kato T Jr, Gotoh Y, Hoffmann A, Ono Y (2008) Negative regulation of constitutive NF-κB and JNK signaling by PKN1–mediated phosphorylation of TRAF1: Genes to Cells. 13: 509–520. - PubMed
-
- Kohno Y, Okaκmoto T, Ishibe T, Nagayama S, Shima Y, et al. (2006) Expression of claudin7 is tightly associated with epithelial structures in synovial sarcomas and regulated by an Ets family transcription factor, ELF3. J Biol Chem 281: 38941–38950. - PubMed
-
- Sapetschnig A, Koch F, Rischitor G, Mennenga T, Suske G (2004) Complexity of translationally controlled transcription factor Sp3 isoform expression. J Biol Chem 279: 42095–42105. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

