Identification of SNP-containing regulatory motifs in the myelodysplastic syndromes model using SNP arrays and gene expression arrays
- PMID: 23327800
- PMCID: PMC3845573
- DOI: 10.5732/cjc.012.10113
Identification of SNP-containing regulatory motifs in the myelodysplastic syndromes model using SNP arrays and gene expression arrays
Abstract
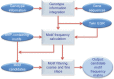
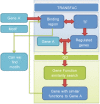
Myelodysplastic syndromes have increased in frequency and incidence in the American population, but patient prognosis has not significantly improved over the last decade. Such improvements could be realized if biomarkers for accurate diagnosis and prognostic stratification were successfully identified. In this study, we propose a method that associates two state-of-the-art array technologies--single nucleotide polymor-phism(SNP) array and gene expression array--with gene motifs considered transcription factor-binding sites (TFBS). We are particularly interested in SNP-containing motifs introduced by genetic variation and mutation as TFBS. The potential regulation of SNP-containing motifs affects only when certain mutations occur. These motifs can be identified from a group of co-expressed genes with copy number variation. Then, we used a sliding window to identify motif candidates near SNPs on gene sequences. The candidates were filtered by coarse thresholding and fine statistical testing. Using the regression-based LARS-EN algorithm and a level-wise sequence combination procedure, we identified 28 SNP-containing motifs as candidate TFBS. We confirmed 21 of the 28 motifs with ChIP-chip fragments in the TRANSFAC database. Another six motifs were validated by TRANSFAC via searching binding fragments on co-regulated genes. The identified motifs and their location genes can be considered potential biomarkers for myelodysplastic syndromes. Thus, our proposed method, a novel strategy for associating two data categories, is capable of integrating information from different sources to identify reliable candidate regulatory SNP-containing motifs introduced by genetic variation and mutation.
Figures




Similar articles
-
Meta-analysis discovery of tissue-specific DNA sequence motifs from mammalian gene expression data.BMC Bioinformatics. 2006 Apr 27;7:229. doi: 10.1186/1471-2105-7-229. BMC Bioinformatics. 2006. PMID: 16643658 Free PMC article.
-
Genome-wide transcription factor binding site/promoter databases for the analysis of gene sets and co-occurrence of transcription factor binding motifs.BMC Genomics. 2010 Mar 1;11:145. doi: 10.1186/1471-2164-11-145. BMC Genomics. 2010. PMID: 20193056 Free PMC article.
-
SNP arrays: comparing diagnostic yields for four platforms in children with developmental delay.BMC Med Genomics. 2014 Dec 24;7:70. doi: 10.1186/s12920-014-0070-0. BMC Med Genomics. 2014. PMID: 25539807 Free PMC article.
-
Significance of genome-wide analysis of copy number alterations and UPD in myelodysplastic syndromes using combined CGH - SNP arrays.Curr Med Chem. 2012;19(22):3739-47. doi: 10.2174/092986712801661121. Curr Med Chem. 2012. PMID: 22680919 Review.
-
Detection of copy number alterations in acute myeloid leukemia and myelodysplastic syndromes.Expert Rev Mol Diagn. 2012 Apr;12(3):253-64. doi: 10.1586/erm.12.18. Expert Rev Mol Diagn. 2012. PMID: 22468816 Review.
Cited by
-
Cancer bioinformatics: detection of chromatin states, SNP-containing motifs, and functional enrichment modules.Chin J Cancer. 2013 Apr;32(4):153-4. doi: 10.5732/cjc.013.10045. Epub 2013 Apr 1. Chin J Cancer. 2013. PMID: 23544450 Free PMC article.
-
Identification and characteristic analysis of enhancers across 13 major cancer types.Precis Clin Med. 2021 Aug 2;4(3):204-208. doi: 10.1093/pcmedi/pbab019. eCollection 2021 Sep. Precis Clin Med. 2021. PMID: 35693214 Free PMC article.
-
Implications of genomic signatures in the differential vulnerability to fetal alcohol exposure in C57BL/6 and DBA/2 mice.Front Genet. 2014 Jun 11;5:173. doi: 10.3389/fgene.2014.00173. eCollection 2014. Front Genet. 2014. PMID: 24966868 Free PMC article.
References
-
- Chen G, Zeng W, Miyazato A, et al. et al. Distinctive gene expression profiles of CD34 cells from patients with myelodysplastic syndrome characterized by specific chromo-somal abnormalities. Blood. 2004;104:4210–4218. - PubMed
-
- Pellagatti A, Esoof N, Watkins F, et al. et al. Gene expression profiling in the myelodysplastic syndromes using cDNA microarray technology. Br J Haematol. 2004;125:576–583. - PubMed
-
- Lastowska M, Viprey V, Santibanez-Koref M, et al. et al. Identification of candidate genes involved in neuroblastoma progression by combining genomic and expression microarrays with survival data. Oncogene. 2007;26:7432–7444. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
