Elevated polar ejection forces stabilize kinetochore-microtubule attachments
- PMID: 23337118
- PMCID: PMC3549975
- DOI: 10.1083/jcb.201211119
Elevated polar ejection forces stabilize kinetochore-microtubule attachments
Abstract
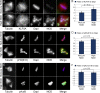
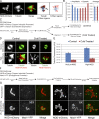
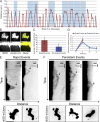
Chromosome biorientation promotes congression and generates tension that stabilizes kinetochore-microtubule (kt-MT) interactions. Forces produced by molecular motors also contribute to chromosome alignment, but their impact on kt-MT attachment stability is unclear. A critical force that acts on chromosomes is the kinesin-10-dependent polar ejection force (PEF). PEFs are proposed to facilitate congression by pushing chromosomes away from spindle poles, although knowledge of the molecular mechanisms underpinning PEF generation is incomplete. Here, we describe a live-cell PEF assay in which tension was applied to chromosomes by manipulating levels of the chromokinesin NOD (no distributive disjunction; Drosophila melanogaster kinesin-10). NOD stabilized syntelic kt-MT attachments in a dose- and motor-dependent manner by overwhelming the ability of Aurora B to mediate error correction. NOD-coated chromatin stretched away from the pole via lateral and end-on interactions with microtubules, and NOD chimeras with either plus end-directed motility or tip-tracking activity produced PEFs. Thus, kt-MT attachment stability is modulated by PEFs, which can be generated by distinct force-producing interactions between chromosomes and dynamic spindle microtubules.
Figures







Similar articles
-
Aurora A Kinase Contributes to a Pole-Based Error Correction Pathway.Curr Biol. 2015 Jul 20;25(14):1842-51. doi: 10.1016/j.cub.2015.06.021. Epub 2015 Jul 9. Curr Biol. 2015. PMID: 26166783 Free PMC article.
-
Dynein prevents erroneous kinetochore-microtubule attachments in mitosis.Cell Cycle. 2015;14(21):3356-61. doi: 10.1080/15384101.2015.1089369. Cell Cycle. 2015. PMID: 26397382 Free PMC article.
-
Insights from an erroneous kinetochore-microtubule attachment state.Bioarchitecture. 2013 May-Jun;3(3):69-76. doi: 10.4161/bioa.25734. Epub 2013 Jul 15. Bioarchitecture. 2013. PMID: 23887229 Free PMC article.
-
Mechanisms of kinesin-7 CENP-E in kinetochore-microtubule capture and chromosome alignment during cell division.Biol Cell. 2019 Jun;111(6):143-160. doi: 10.1111/boc.201800082. Epub 2019 Feb 26. Biol Cell. 2019. PMID: 30784092 Review.
-
Regulation of kinetochore-microtubule attachments by Aurora B kinase.Biochem Soc Trans. 2009 Oct;37(Pt 5):976-80. doi: 10.1042/BST0370976. Biochem Soc Trans. 2009. PMID: 19754435 Review.
Cited by
-
Role of spatial patterns and kinetochore architecture in spindle morphogenesis.Semin Cell Dev Biol. 2021 Sep;117:75-85. doi: 10.1016/j.semcdb.2021.03.016. Epub 2021 Apr 6. Semin Cell Dev Biol. 2021. PMID: 33836948 Free PMC article. Review.
-
Mammalian kinetochores count attached microtubules in a sensitive and switch-like manner.J Cell Biol. 2019 Nov 4;218(11):3583-3596. doi: 10.1083/jcb.201902105. Epub 2019 Sep 6. J Cell Biol. 2019. PMID: 31492713 Free PMC article.
-
Mechanisms of Chromosome Congression during Mitosis.Biology (Basel). 2017 Feb 17;6(1):13. doi: 10.3390/biology6010013. Biology (Basel). 2017. PMID: 28218637 Free PMC article. Review.
-
Chromosomal instability: Stretching the role of checkpoint silencing.Curr Biol. 2021 Apr 26;31(8):R386-R389. doi: 10.1016/j.cub.2021.02.014. Curr Biol. 2021. PMID: 33905696 Free PMC article.
-
The Tubulin Code: A Navigation System for Chromosomes during Mitosis.Trends Cell Biol. 2016 Oct;26(10):766-775. doi: 10.1016/j.tcb.2016.06.001. Epub 2016 Jun 22. Trends Cell Biol. 2016. PMID: 27344407 Free PMC article. Review.
References
-
- Adams R.R., Maiato H., Earnshaw W.C., Carmena M. 2001. Essential roles of Drosophila inner centromere protein (INCENP) and aurora B in histone H3 phosphorylation, metaphase chromosome alignment, kinetochore disjunction, and chromosome segregation. J. Cell Biol. 153:865–880 10.1083/jcb.153.4.865 - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

