Evolutionary conservation of the polyproline II conformation surrounding intrinsically disordered phosphorylation sites
- PMID: 23341186
- PMCID: PMC3610046
- DOI: 10.1002/pro.2217
Evolutionary conservation of the polyproline II conformation surrounding intrinsically disordered phosphorylation sites
Abstract
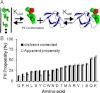
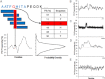
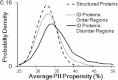
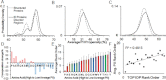
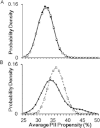
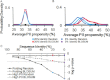
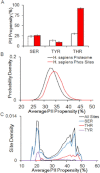
Intrinsically disordered (ID) proteins function in the absence of a unique stable structure and appear to challenge the classic structure-function paradigm. The extent to which ID proteins take advantage of subtle conformational biases to perform functions, and whether signals for such mechanism can be identified in proteome-wide studies is not well understood. Of particular interest is the polyproline II (PII) conformation, suggested to be highly populated in unfolded proteins. We experimentally determine a complete calorimetric propensity scale for the PII conformation. Projection of the scale into representative eukaryotic proteomes reveals significant PII bias in regions coding for ID proteins. Importantly, enrichment of PII in ID proteins, or protein segments, is also captured by other PII scales, indicating that this enrichment is robustly encoded and universally detectable regardless of the method of PII propensity determination. Gene ontology (GO) terms obtained using our PII scale and other scales demonstrate a consensus for molecular functions performed by high PII proteins across the proteome. Perhaps the most striking result of the GO analysis is conserved enrichment (P < 10(-8) ) of phosphorylation sites in high PII regions found by all PII scales. Subsequent conformational analysis reveals a phosphorylation-dependent modulation of PII, suggestive of a conserved "tunability" within these regions. In summary, the application of an experimentally determined polyproline II (PII) propensity scale to proteome-wide sequence analysis and gene ontology reveals an enrichment of PII bias near disordered phosphorylation sites that is conserved throughout eukaryotes.
Copyright © 2013 The Protein Society.
Figures


Similar articles
-
Temperature and urea have opposing impacts on polyproline II conformational bias.Biochemistry. 2013 Feb 5;52(5):949-58. doi: 10.1021/bi301435p. Epub 2013 Jan 27. Biochemistry. 2013. PMID: 23350874 Free PMC article.
-
Hidden dynamic signatures drive substrate selectivity in the disordered phosphoproteome.Proc Natl Acad Sci U S A. 2020 Sep 22;117(38):23606-23616. doi: 10.1073/pnas.1921473117. Epub 2020 Sep 8. Proc Natl Acad Sci U S A. 2020. PMID: 32900925 Free PMC article.
-
Proteome-wide discovery of evolutionary conserved sequences in disordered regions.Sci Signal. 2012 Mar 13;5(215):rs1. doi: 10.1126/scisignal.2002515. Sci Signal. 2012. PMID: 22416277 Free PMC article.
-
The relevance of short peptides for an understanding of unfolded and intrinsically disordered proteins.Phys Chem Chem Phys. 2023 May 3;25(17):11908-11933. doi: 10.1039/d3cp00483j. Phys Chem Chem Phys. 2023. PMID: 37096579 Review.
-
Conformational propensities and residual structures in unfolded peptides and proteins.Mol Biosyst. 2012 Jan;8(1):122-33. doi: 10.1039/c1mb05225j. Epub 2011 Aug 30. Mol Biosyst. 2012. PMID: 21879108 Review.
Cited by
-
Impact of Heat on Coil Hydrodynamic Size Yields the Energetics of Denatured State Conformational Bias.J Phys Chem B. 2019 Nov 27;123(47):10014-10024. doi: 10.1021/acs.jpcb.9b09088. Epub 2019 Nov 14. J Phys Chem B. 2019. PMID: 31679343 Free PMC article.
-
An Unbound Proline-Rich Signaling Peptide Frequently Samples Cis Conformations in Gaussian Accelerated Molecular Dynamics Simulations.Front Mol Biosci. 2021 Nov 15;8:734169. doi: 10.3389/fmolb.2021.734169. eCollection 2021. Front Mol Biosci. 2021. PMID: 34869581 Free PMC article.
-
Convergent behavior of extended stalk regions from staphylococcal surface proteins with widely divergent sequence patterns.Protein Sci. 2023 Aug;32(8):e4707. doi: 10.1002/pro.4707. Protein Sci. 2023. PMID: 37334491 Free PMC article.
-
Structural and Energetic Characterization of the Denatured State from the Perspectives of Peptides, the Coil Library, and Intrinsically Disordered Proteins.Molecules. 2021 Jan 26;26(3):634. doi: 10.3390/molecules26030634. Molecules. 2021. PMID: 33530506 Free PMC article. Review.
-
Convergent behavior of extended stalk regions from staphylococcal surface proteins with widely divergent sequence patterns.bioRxiv [Preprint]. 2023 Jan 7:2023.01.06.523059. doi: 10.1101/2023.01.06.523059. bioRxiv. 2023. Update in: Protein Sci. 2023 Aug;32(8):e4707. doi: 10.1002/pro.4707. PMID: 36711672 Free PMC article. Updated. Preprint.
References
-
- Ward J, Sodhi J, McGuffin L, Buxton B, Jones D. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol. 2004;337:635–645. - PubMed
-
- Oldfield C, Cheng Y, Cortese M, Brown C, Uversky V, Dunker A. Comparing and combining predictors of mostly disordered proteins. Biochemistry. 2005;44:1989–2000. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

