Reassessing the type I dehydroquinate dehydratase catalytic triad: kinetic and structural studies of Glu86 mutants
- PMID: 23341204
- PMCID: PMC3610047
- DOI: 10.1002/pro.2218
Reassessing the type I dehydroquinate dehydratase catalytic triad: kinetic and structural studies of Glu86 mutants
Abstract
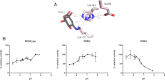
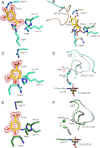
Dehydroquinate dehydratase (DHQD) catalyzes the third reaction in the biosynthetic shikimate pathway. Type I DHQDs are members of the greater aldolase superfamily, a group of enzymes that contain an active site lysine that forms a Schiff base intermediate. Three residues (Glu86, His143, and Lys170 in the Salmonella enterica DHQD) have previously been proposed to form a triad vital for catalysis. While the roles of Lys170 and His143 are well defined-Lys170 forms the Schiff base with the substrate and His143 shuttles protons in multiple steps in the reaction-the role of Glu86 remains poorly characterized. To probe Glu86's role, Glu86 mutants were generated and subjected to biochemical and structural study. The studies presented here demonstrate that mutant enzymes retain catalytic proficiency, calling into question the previously attributed role of Glu86 in catalysis and suggesting that His143 and Lys170 function as a catalytic dyad. Structures of the Glu86Ala (E86A) mutant in complex with covalently bound reaction intermediate reveal a conformational change of the His143 side chain. This indicates a predominant steric role for Glu86, to maintain the His143 side chain in position consistent with catalysis. The structures also explain why the E86A mutant is optimally active at more acidic conditions than the wild-type enzyme. In addition, a complex with the reaction product reveals a novel, likely nonproductive, binding mode that suggests a mechanism of competitive product inhibition and a potential strategy for the design of therapeutics.
Copyright © 2013 The Protein Society.
Figures
Similar articles
-
Crystal structures of type I dehydroquinate dehydratase in complex with quinate and shikimate suggest a novel mechanism of Schiff base formation.Biochemistry. 2014 Feb 11;53(5):872-80. doi: 10.1021/bi4015506. Epub 2014 Jan 31. Biochemistry. 2014. PMID: 24437575 Free PMC article.
-
New insights into the mechanism of the Schiff base hydrolysis catalyzed by type I dehydroquinate dehydratase from S. enterica: a theoretical study.Org Biomol Chem. 2012 Sep 21;10(35):7037-44. doi: 10.1039/c2ob25605c. Epub 2012 Jul 31. Org Biomol Chem. 2012. PMID: 22847490
-
Insights into the mechanism of type I dehydroquinate dehydratases from structures of reaction intermediates.J Biol Chem. 2011 Feb 4;286(5):3531-9. doi: 10.1074/jbc.M110.192831. Epub 2010 Nov 18. J Biol Chem. 2011. PMID: 21087925 Free PMC article.
-
A conserved surface loop in type I dehydroquinate dehydratases positions an active site arginine and functions in substrate binding.Biochemistry. 2011 Mar 29;50(12):2357-63. doi: 10.1021/bi102020s. Epub 2011 Feb 21. Biochemistry. 2011. PMID: 21291284 Free PMC article.
-
Experiences with the shikimate-pathway enzymes as targets for rational drug design.Biochem Soc Trans. 2003 Jun;31(Pt 3):548-52. doi: 10.1042/bst0310548. Biochem Soc Trans. 2003. PMID: 12773154 Review.
Cited by
-
Structural and functional analysis of betaine aldehyde dehydrogenase from Staphylococcus aureus.Acta Crystallogr D Biol Crystallogr. 2015 May;71(Pt 5):1159-75. doi: 10.1107/S1399004715004228. Epub 2015 Apr 25. Acta Crystallogr D Biol Crystallogr. 2015. PMID: 25945581 Free PMC article.
-
Structure of type II dehydroquinase from Pseudomonas aeruginosa.Acta Crystallogr F Struct Biol Commun. 2014 Nov;70(Pt 11):1485-91. doi: 10.1107/S2053230X14020214. Epub 2014 Oct 25. Acta Crystallogr F Struct Biol Commun. 2014. PMID: 25372814 Free PMC article.
-
Molecular analysis and essentiality of Aro1 shikimate biosynthesis multi-enzyme in Candida albicans.Life Sci Alliance. 2022 May 5;5(8):e202101358. doi: 10.26508/lsa.202101358. Print 2022 Aug. Life Sci Alliance. 2022. PMID: 35512834 Free PMC article.
-
Crystal structures of type I dehydroquinate dehydratase in complex with quinate and shikimate suggest a novel mechanism of Schiff base formation.Biochemistry. 2014 Feb 11;53(5):872-80. doi: 10.1021/bi4015506. Epub 2014 Jan 31. Biochemistry. 2014. PMID: 24437575 Free PMC article.
References
-
- Gourley DG, Shrive AK, Polikarpov I, Krell T, Coggins JR, Hawkins AR, Isaacs NW, Sawyer L. The two types of 3-dehydroquinase have distinct structures but catalyze the same overall reaction. Nat Struct Biol. 1999;6:521–525. - PubMed
-
- Choi KH, Lai V, Foster CE, Morris AJ, Tolan DR, Allen KN. New superfamily members identified for Schiff-base enzymes based on verification of catalytically essential residues. Biochemistry. 2006;45:8546–8555. - PubMed
-
- Butler JR, Alworth WL, Nugent MJ. Mechanism of dehydroquinase catalyzed dehydration. I. Formation of a Schiff-base intermediate. J Am Chem Soc. 1974;96:1617–1618.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

