Organ evolution in angiosperms driven by correlated divergences of gene sequences and expression patterns
- PMID: 23341336
- PMCID: PMC3584551
- DOI: 10.1105/tpc.112.106716
Organ evolution in angiosperms driven by correlated divergences of gene sequences and expression patterns
Abstract
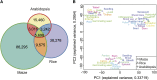
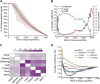
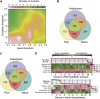
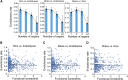
The evolution of a species involves changes in its genome and its transcriptome. Divergence in expression patterns may be more important than divergence in sequences for determining phenotypic changes, particularly among closely related species. We examined the relationships between organ evolution, sequence evolution, and expression evolution in Arabidopsis thaliana, rice (Oryza sativa), and maize (Zea mays). We found correlated divergence of gene sequences and expression patterns, with distinct divergence rates that depend on the organ types in which a gene is expressed. For instance, genes specifically expressed in reproductive organs (i.e., stamen) evolve more quickly than those specifically expressed in vegetative organs (e.g., root). The different rates in organ evolution may be due to different degrees of functional constraint associated with the different physiological functions of plant organs. Additionally, the evolutionary rate of a gene sequence is correlated with the breadth of its expression in terms of the number of tissues, the number of coregulation modules, and the number of species in which the gene is expressed, as well as the number of genes with which it may interact. This linkage supports the hypothesis that constitutively expressed genes may experience higher levels of functional constraint accumulated from multiple tissues than do tissue-specific genes.
Figures


Comment in
-
Gene expression in angiosperm organ evolution.Plant Cell. 2013 Feb;25(2):357. doi: 10.1105/tpc.113.250210. Epub 2013 Feb 12. Plant Cell. 2013. PMID: 23404885 Free PMC article. No abstract available.
References
-
- Bergmann, S., Ihmels, J., and Barkai, N. (2003). Iterative signature algorithm for the analysis of large-scale gene expression data. Phys. Rev. E Stat Nonlin. Soft Matter Phys. 67: 031902. - PubMed
-
- Brawand D., et al. (2011). The evolution of gene expression levels in mammalian organs. Nature 478: 343–348 - PubMed
-
- Cao J., et al. (2011). Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat. Genet. 43: 956–963 - PubMed
-
- Clark N.L., Aagaard J.E., Swanson W.J. (2006). Evolution of reproductive proteins from animals and plants. Reproduction 131: 11–22 - PubMed
-
- Csárdi G., Kutalik Z., Bergmann S. (2010). Modular analysis of gene expression data with R. Bioinformatics 26: 1376–1377 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

