Down-regulation of vitamin D receptor in mammospheres: implications for vitamin D resistance in breast cancer and potential for combination therapy
- PMID: 23341935
- PMCID: PMC3544824
- DOI: 10.1371/journal.pone.0053287
Down-regulation of vitamin D receptor in mammospheres: implications for vitamin D resistance in breast cancer and potential for combination therapy
Erratum in
- PLoS One. 2013;8(10). doi:10.1371/annotation/5326d117-3f31-4e43-a5c4-9e1fb41719e9
Abstract
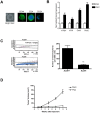
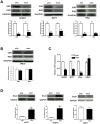
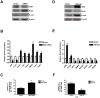
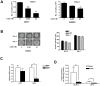
Vitamin D signaling in mammary cancer stem cells (MCSCs), which are implicated in the initiation and progression of breast cancer, is poorly understood. In this study, we examined vitamin D signaling in mammospheres which are enriched in MCSCs from established breast cancer cell lines. Breast cancer cells positive for aldehyde dehydrogenase (ALDH(+)) had increased ability to form mammospheres compared to ALDH(-) cells. These mammospheres expressed MCSC-specific markers and generated transplantable xenografts in nude mice. Vitamin D receptor (VDR) was significantly down-regulated in mammospheres, as well as in ALDH(+) breast cancer cells. TN aggressive human breast tumors as well as transplantable xenografts obtained from SKBR3 expressed significantly lower levels of VDR but higher levels of CD44 expression. Snail was up-regulated in mammospheres isolated from breast cancer cells. Inhibition of VDR expression by siRNA led to a significant change in key EMT-specific transcription factors and increased the ability of these cells to form mammospheres. On the other hand, over-expression of VDR led to a down-regulation of Snail but increased expression of E-cad and significantly compromised the ability of cells to form mammospheres. Mammospheres were relatively insensitive to treatment with 1,25-dihydroxyvitamin D (1,25D), the active form of vitamin D, compared to more differentiated cancer cells grown in presence of serum. Treatment of H-Ras transformed HMLE(HRas) cells with DETA NONOate, a nitric oxide (NO)-donor led to induction of MAP-kinase phosphatase -1 (MKP-1) and dephosphorylation of ERK1/2 in the mammospheres. Combined treatment of these cells with 1,25D and a low-concentration of DETA NONOate led to a significant decrease in the overall size of mammospheres and reduced tumor volume in nude mice. Our findings therefore, suggest that combination therapy using 1,25D with drugs specifically targeting key survival pathways in MCSCs warrant testing in prospective clinical trial for treatment of aggressive breast cancer.
Conflict of interest statement
Figures




Similar articles
-
PARP3 controls TGFβ and ROS driven epithelial-to-mesenchymal transition and stemness by stimulating a TG2-Snail-E-cadherin axis.Oncotarget. 2016 Sep 27;7(39):64109-64123. doi: 10.18632/oncotarget.11627. Oncotarget. 2016. PMID: 27579892 Free PMC article.
-
Combination therapy with epigenetic-targeted and chemotherapeutic drugs delivered by nanoparticles to enhance the chemotherapy response and overcome resistance by breast cancer stem cells.J Control Release. 2015 May 10;205:7-14. doi: 10.1016/j.jconrel.2014.11.011. Epub 2014 Nov 15. J Control Release. 2015. PMID: 25445694
-
SNAIL regulates interleukin-8 expression, stem cell-like activity, and tumorigenicity of human colorectal carcinoma cells.Gastroenterology. 2011 Jul;141(1):279-91, 291.e1-5. doi: 10.1053/j.gastro.2011.04.008. Epub 2011 Apr 16. Gastroenterology. 2011. PMID: 21640118
-
Mammary stem cells and breast cancer--role of Notch signalling.Stem Cell Rev. 2007 Jun;3(2):169-75. doi: 10.1007/s12015-007-0023-5. Stem Cell Rev. 2007. PMID: 17873349 Review.
-
Targets of vitamin D receptor signaling in the mammary gland.J Bone Miner Res. 2007 Dec;22 Suppl 2:V86-90. doi: 10.1359/jbmr.07s204. J Bone Miner Res. 2007. PMID: 18290729 Review.
Cited by
-
Dynamic interplay of nuclear receptors in tumor cell plasticity and drug resistance: Shifting gears in malignant transformations and applications in cancer therapeutics.Cancer Metastasis Rev. 2024 Mar;43(1):321-362. doi: 10.1007/s10555-024-10171-0. Epub 2024 Mar 22. Cancer Metastasis Rev. 2024. PMID: 38517618 Review.
-
The role of vitamin D in reducing cancer risk and progression.Nat Rev Cancer. 2014 May;14(5):342-57. doi: 10.1038/nrc3691. Epub 2014 Apr 4. Nat Rev Cancer. 2014. PMID: 24705652 Review.
-
Tumor Expression of Vitamin D Receptor and Breast Cancer Histopathological Characteristics and Prognosis.Clin Cancer Res. 2017 Jan 1;23(1):97-103. doi: 10.1158/1078-0432.CCR-16-0075. Epub 2016 Jul 12. Clin Cancer Res. 2017. PMID: 27407090 Free PMC article.
-
Vitamin D3 pretreatment regulates renal inflammatory responses during lipopolysaccharide-induced acute kidney injury.Sci Rep. 2015 Dec 22;5:18687. doi: 10.1038/srep18687. Sci Rep. 2015. PMID: 26691774 Free PMC article.
-
Role of nuclear receptors in breast cancer stem cells.World J Stem Cells. 2016 Mar 26;8(3):62-72. doi: 10.4252/wjsc.v8.i3.62. World J Stem Cells. 2016. PMID: 27022437 Free PMC article. Review.
References
-
- Toner CD, Davis CD, Milner JA (2010) The vitamin D and cancer conundrum: aiming at a moving target J AM DIET ASSOC. 110: 1492–1500. - PubMed
-
- Pérez-López FR, Chedraui P, Haya J (2009) Review article: vitamin D acquisition and breast cancer risk. Reprod Sci 16: 7–19. - PubMed
-
- McKay JD, McCullough ML, Ziegler RG, Kraft P, Saltzman BS, et al. (2009) Vitamin D receptor polymorphisms and breast cancer risk: results from the National Cancer Institute Breast and Prostate Cancer Cohort Consortium. Cancer Epidemiol Biomarkers Prev 18: 297–305. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

