Proteomic profiling of Mycobacterium tuberculosis identifies nutrient-starvation-responsive toxin-antitoxin systems
- PMID: 23345537
- PMCID: PMC3650330
- DOI: 10.1074/mcp.M112.018846
Proteomic profiling of Mycobacterium tuberculosis identifies nutrient-starvation-responsive toxin-antitoxin systems
Abstract
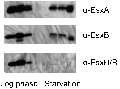
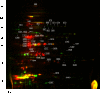
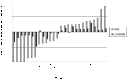
In order to successfully enter the latent stage, Mycobacterium tuberculosis must adapt to conditions such as nutrient limitation and hypoxia. In vitro models that mimic latent infection are valuable tools for describing the changes in metabolism that occur when the bacterium exists in a non-growing form. We used two complementary proteomic approaches, label-free LC-MS/MS analysis and two-dimensional difference gel electrophoresis, to determine the proteome profile of extracellular proteins from M. tuberculosis cultured under nutrient starvation. Through the label-free LC-MS/MS analysis of fractionated samples, 1176 proteins were identified from culture filtrates of log phase and nutrient-starved cultures, and the protein levels of 230 proteins were increased in nutrient-starved culture filtrates, whereas those of 208 proteins were decreased. By means of Gene Ontology clustering analysis, significant differences in the overall metabolism during nutrient starvation were detected. Notably, members of the toxin-antitoxin systems were present in larger quantities in nutrient-starved cultures, supporting a role for these global modules as M. tuberculosis switches its metabolism into dormancy. Decreased abundance of proteins involved in amino acid and protein synthesis was apparent, as well as changes in the lipid metabolism. Further analysis of the dataset identified increased abundance of lipoproteins and decreased abundance of ESAT-6 family proteins. Results from the two-dimensional difference gel electrophoresis proteomics demonstrated overall agreement with the LC-MS/MS data and added complementary insights about protein degradation and modification.
Figures
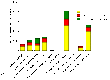
References
-
- Garton N. J., Waddell S. J., Sherratt A. L., Lee S. M., Smith R. J., Senner C., Hinds J., Rajakumar K., Adegbola R. A., Besra G. S., Butcher P. D., Barer M. R. (2008) Cytological and transcript analyses reveal fat and lazy persister-like bacilli in tuberculous sputum. PLoS Med. 5, e75. - PMC - PubMed
-
- Betts J. C., Lukey P. T., Robb L. C., McAdam R. A., Duncan K. (2002) Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence by gene and protein expression profiling. Mol. Microbiol. 43, 717–731 - PubMed
-
- Andersen P., Munk M. E., Pollock J. M., Doherty T. M. (2000) Specific immune-based diagnosis of tuberculosis. Lancet 356, 1099–1104 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

