Insight into centromere-binding properties of ParB proteins: a secondary binding motif is essential for bacterial genome maintenance
- PMID: 23345617
- PMCID: PMC3597684
- DOI: 10.1093/nar/gkt018
Insight into centromere-binding properties of ParB proteins: a secondary binding motif is essential for bacterial genome maintenance
Abstract
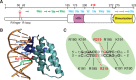
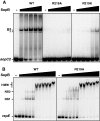
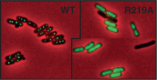
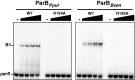
ParB proteins are one of the three essential components of partition systems that actively segregate bacterial chromosomes and plasmids. In binding to centromere sequences, ParB assembles as nucleoprotein structures called partition complexes. These assemblies are the substrates for the partitioning process that ensures DNA molecules are segregated to both sides of the cell. We recently identified the sopC centromere nucleotides required for binding to the ParB homologue of plasmid F, SopB. This analysis also suggested a role in sopC binding for an arginine residue, R219, located outside the helix-turn-helix (HTH) DNA-binding motif previously shown to be the only determinant for sopC-specific binding. Here, we demonstrated that the R219 residue is critical for SopB binding to sopC during partition. Mutating R219 to alanine or lysine abolished partition by preventing partition complex assembly. Thus, specificity of SopB binding relies on two distinct motifs, an HTH and an arginine residue, which define a split DNA-binding domain larger than previously thought. Bioinformatic analysis over a broad range of chromosomal ParBs generalized our findings with the identification of a non-HTH positively charged residue essential for partition and centromere binding, present in a newly identified highly conserved motif. We propose that ParB proteins possess two DNA-binding motifs that form an extended centromere-binding domain, providing high specificity.
Figures


Similar articles
-
Structures of ParB bound to DNA reveal mechanism of partition complex formation.Nature. 2005 Nov 24;438(7067):516-9. doi: 10.1038/nature04149. Nature. 2005. PMID: 16306995
-
Molecular analysis of the pRA2 partitioning region: ParB autoregulates parAB transcription and forms a nucleoprotein complex with the plasmid partition site, parS.Mol Microbiol. 2001 May;40(3):621-33. doi: 10.1046/j.1365-2958.2001.02405.x. Mol Microbiol. 2001. PMID: 11359568
-
Centromere binding specificity in assembly of the F plasmid partition complex.Nucleic Acids Res. 2011 Sep 1;39(17):7477-86. doi: 10.1093/nar/gkr457. Epub 2011 Jun 7. Nucleic Acids Res. 2011. PMID: 21653553 Free PMC article.
-
ParB Partition Proteins: Complex Formation and Spreading at Bacterial and Plasmid Centromeres.Front Mol Biosci. 2016 Aug 29;3:44. doi: 10.3389/fmolb.2016.00044. eCollection 2016. Front Mol Biosci. 2016. PMID: 27622187 Free PMC article. Review.
-
Structural biology of plasmid partition: uncovering the molecular mechanisms of DNA segregation.Biochem J. 2008 May 15;412(1):1-18. doi: 10.1042/BJ20080359. Biochem J. 2008. PMID: 18426389 Review.
Cited by
-
A conserved mechanism drives partition complex assembly on bacterial chromosomes and plasmids.Mol Syst Biol. 2018 Nov 16;14(11):e8516. doi: 10.15252/msb.20188516. Mol Syst Biol. 2018. PMID: 30446599 Free PMC article.
-
Defining the role of ATP hydrolysis in mitotic segregation of bacterial plasmids.PLoS Genet. 2013;9(12):e1003956. doi: 10.1371/journal.pgen.1003956. Epub 2013 Dec 19. PLoS Genet. 2013. PMID: 24367270 Free PMC article.
-
DNA Segregation in Enterobacteria.EcoSal Plus. 2023 Dec 12;11(1):eesp00382020. doi: 10.1128/ecosalplus.esp-0038-2020. Epub 2023 May 9. EcoSal Plus. 2023. PMID: 37220081 Free PMC article. Review.
-
Segrosome Complex Formation during DNA Trafficking in Bacterial Cell Division.Front Mol Biosci. 2016 Sep 9;3:51. doi: 10.3389/fmolb.2016.00051. eCollection 2016. Front Mol Biosci. 2016. PMID: 27668216 Free PMC article. Review.
-
Replication and Active Partition of Integrative and Conjugative Elements (ICEs) of the SXT/R391 Family: The Line between ICEs and Conjugative Plasmids Is Getting Thinner.PLoS Genet. 2015 Jun 10;11(6):e1005298. doi: 10.1371/journal.pgen.1005298. eCollection 2015 Jun. PLoS Genet. 2015. PMID: 26061412 Free PMC article.
References
-
- Salje J. Plasmid segregation: how to survive as an extra piece of DNA. Crit. Rev. Biochem. Mol. Biol. 2010;45:296–317. - PubMed
-
- Gerdes K, Moller-Jensen J, Bugge Jensen R. Plasmid and chromosome partitioning: surprises from phylogeny. Mol. Microbiol. 2000;37:455–466. - PubMed
-
- Marston AL, Errington J. Dynamic movement of the ParA-like Soj protein of B. subtilis and its dual role in nucleoid organization and developmental regulation. Mol. Cell. 1999;4:673–682. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

