Development of bis-locked nucleic acid (bisLNA) oligonucleotides for efficient invasion of supercoiled duplex DNA
- PMID: 23345620
- PMCID: PMC3597675
- DOI: 10.1093/nar/gkt007
Development of bis-locked nucleic acid (bisLNA) oligonucleotides for efficient invasion of supercoiled duplex DNA
Abstract
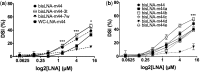
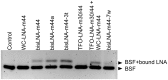
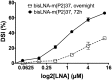
In spite of the many developments in synthetic oligonucleotide (ON) chemistry and design, invasion into double-stranded DNA (DSI) under physiological salt and pH conditions remains a challenge. In this work, we provide a new ON tool based on locked nucleic acids (LNAs), designed for strand invasion into duplex DNA (DSI). We thus report on the development of a clamp type of LNA ON-bisLNA-with capacity to bind and invade into supercoiled double-stranded DNA. The bisLNA links a triplex-forming, Hoogsteen-binding, targeting arm with a strand-invading Watson-Crick binding arm. Optimization was carried out by varying the number and location of LNA nucleotides and the length of the triplex-forming versus strand-invading arms. Single-strand regions in target duplex DNA were mapped using chemical probing. By combining design and increase in LNA content, it was possible to achieve a 100-fold increase in potency with 30% DSI at 450 nM using a bisLNA to plasmid ratio of only 21:1. Although this first conceptual report does not address the utility of bisLNA for the targeting of DNA in a chromosomal context, it shows bisLNA as a promising candidate for interfering also with cellular genes.
Figures





Similar articles
-
Next-generation bis-locked nucleic acids with stacking linker and 2'-glycylamino-LNA show enhanced DNA invasion into supercoiled duplexes.Nucleic Acids Res. 2016 Mar 18;44(5):2007-19. doi: 10.1093/nar/gkw021. Epub 2016 Feb 8. Nucleic Acids Res. 2016. PMID: 26857548 Free PMC article.
-
The ability of locked nucleic acid oligonucleotides to pre-structure the double helix: A molecular simulation and binding study.PLoS One. 2019 Feb 12;14(2):e0211651. doi: 10.1371/journal.pone.0211651. eCollection 2019. PLoS One. 2019. PMID: 30753192 Free PMC article.
-
Use of locked nucleic acid oligonucleotides to add functionality to plasmid DNA.Nucleic Acids Res. 2003 Oct 15;31(20):5817-30. doi: 10.1093/nar/gkg801. Nucleic Acids Res. 2003. PMID: 14530430 Free PMC article.
-
Optimizing anti-gene oligonucleotide 'Zorro-LNA' for improved strand invasion into duplex DNA.Nucleic Acids Res. 2011 Feb;39(3):1142-54. doi: 10.1093/nar/gkq835. Epub 2010 Sep 21. Nucleic Acids Res. 2011. PMID: 20860997 Free PMC article.
-
Targeting DNA with triplex-forming oligonucleotides to modify gene sequence.Biochimie. 2008 Aug;90(8):1109-16. doi: 10.1016/j.biochi.2008.04.004. Epub 2008 Apr 18. Biochimie. 2008. PMID: 18460344 Review.
Cited by
-
Anti-gene oligonucleotides targeting Friedreich's ataxia expanded GAA⋅TTC repeats increase Frataxin expression.Mol Ther Nucleic Acids. 2025 Apr 17;36(2):102541. doi: 10.1016/j.omtn.2025.102541. eCollection 2025 Jun 10. Mol Ther Nucleic Acids. 2025. PMID: 40487352 Free PMC article.
-
CTG repeat-targeting oligonucleotides for down-regulating Huntingtin expression.Nucleic Acids Res. 2017 May 19;45(9):5153-5169. doi: 10.1093/nar/gkx111. Nucleic Acids Res. 2017. PMID: 28334749 Free PMC article.
-
Hydrogel-Assisted Antisense LNA Gapmer Delivery for In Situ Gene Silencing in Spinal Cord Injury.Mol Ther Nucleic Acids. 2018 Jun 1;11:393-406. doi: 10.1016/j.omtn.2018.03.009. Epub 2018 Mar 20. Mol Ther Nucleic Acids. 2018. PMID: 29858074 Free PMC article.
-
Next-generation bis-locked nucleic acids with stacking linker and 2'-glycylamino-LNA show enhanced DNA invasion into supercoiled duplexes.Nucleic Acids Res. 2016 Mar 18;44(5):2007-19. doi: 10.1093/nar/gkw021. Epub 2016 Feb 8. Nucleic Acids Res. 2016. PMID: 26857548 Free PMC article.
-
Synthesis and characterization of oligodeoxyribonucleotides modified with 2'-thio-2'-deoxy-2'-S-(pyren-1-yl)methyluridine.Bioorg Med Chem Lett. 2015 Sep 15;25(18):3999-4004. doi: 10.1016/j.bmcl.2015.07.002. Epub 2015 Jul 7. Bioorg Med Chem Lett. 2015. PMID: 26254942 Free PMC article.
References
-
- Bennett CF, Swayze EE. RNA targeting therapeutics: molecular mechanisms of antisense oligonucleotides as a therapeutic platform. Annu. Rev. Pharmacol. Toxicol. 2010;50:259–293. - PubMed
-
- Moser HE, Dervan PB. Sequence-specific cleavage of double helical DNA by triple helix formation. Science. 1987;238:645–650. - PubMed
-
- Le Doan T, Perrouault L, Praseuth D, Habhoub N, Decout JL, Thuong NT, Lhomme J, Helene C. Sequence-specific recognition, photocrosslinking and cleavage of the DNA double helix by an oligo-[alpha]-thymidylate covalently linked to an azidoproflavine derivative. Nucleic Acids Res. 1987;15:7749–7760. - PMC - PubMed
-
- Kim HG, Miller DM. Inhibition of in vitro transcription by a triplex-forming oligonucleotide targeted to human c-myc P2 promoter. Biochemistry. 1995;34:8165–8171. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

