Application of whole exome sequencing to identify disease-causing variants in inherited human diseases
- PMID: 23346032
- PMCID: PMC3543920
- DOI: 10.5808/GI.2012.10.4.214
Application of whole exome sequencing to identify disease-causing variants in inherited human diseases
Abstract
The recent advent of next-generation sequencing technologies has dramatically changed the nature of biomedical research. Human genetics is no exception-it has never been easier to interrogate human patient genomes at the nucleotide level to identify disease-associated variants. To further facilitate the efficiency of this approach, whole exome sequencing (WES) was first developed in 2009. Over the past three years, multiple groups have demonstrated the power of WES through robust disease-associated variant discoveries across a diverse spectrum of human diseases. Here, we review the application of WES to different types of inherited human diseases and discuss analytical challenges and possible solutions, with the aim of providing a practical guide for the effective use of this technology.
Keywords: discovery of disease-causing variants; inherited human disease; next-generation sequencing; whole exome sequencing.
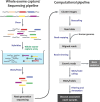
Figures


Similar articles
-
Recent Advances in Autism Spectrum Disorders: Applications of Whole Exome Sequencing Technology.Psychiatry Investig. 2016 May;13(3):255-64. doi: 10.4306/pi.2016.13.3.255. Epub 2016 May 18. Psychiatry Investig. 2016. PMID: 27247591 Free PMC article. Review.
-
Whole exome sequencing: Uncovering causal genetic variants for ocular diseases.Exp Eye Res. 2017 Nov;164:139-150. doi: 10.1016/j.exer.2017.08.013. Epub 2017 Aug 24. Exp Eye Res. 2017. PMID: 28844620 Review.
-
Whole-genome sequencing is more powerful than whole-exome sequencing for detecting exome variants.Proc Natl Acad Sci U S A. 2015 Apr 28;112(17):5473-8. doi: 10.1073/pnas.1418631112. Epub 2015 Mar 31. Proc Natl Acad Sci U S A. 2015. PMID: 25827230 Free PMC article.
-
Candidate gene discovery in autoimmunity by using extreme phenotypes, next generation sequencing and whole exome capture.Autoimmun Rev. 2015 Mar;14(3):204-9. doi: 10.1016/j.autrev.2014.10.021. Epub 2014 Nov 1. Autoimmun Rev. 2015. PMID: 25447288 Review.
-
Inconsistency and features of single nucleotide variants detected in whole exome sequencing versus transcriptome sequencing: A case study in lung cancer.Methods. 2015 Jul 15;83:118-27. doi: 10.1016/j.ymeth.2015.04.016. Epub 2015 Apr 23. Methods. 2015. PMID: 25913717 Free PMC article.
Cited by
-
The Landscape of Point Mutations in Human Protein Coding Genes Leading to Pregnancy Loss.Int J Mol Sci. 2023 Dec 17;24(24):17572. doi: 10.3390/ijms242417572. Int J Mol Sci. 2023. PMID: 38139401 Free PMC article. Review.
-
Targeted sequencing of large genomic regions with CATCH-Seq.PLoS One. 2014 Oct 30;9(10):e111756. doi: 10.1371/journal.pone.0111756. eCollection 2014. PLoS One. 2014. PMID: 25357200 Free PMC article.
-
SECNVs: A Simulator of Copy Number Variants and Whole-Exome Sequences From Reference Genomes.Front Genet. 2020 Feb 21;11:82. doi: 10.3389/fgene.2020.00082. eCollection 2020. Front Genet. 2020. PMID: 32153642 Free PMC article.
-
A Mutation in ZNF143 as a Novel Candidate Gene for Endothelial Corneal Dystrophy.J Clin Med. 2019 Aug 6;8(8):1174. doi: 10.3390/jcm8081174. J Clin Med. 2019. PMID: 31390831 Free PMC article.
-
Rare cases of congenital arthrogryposis multiplex caused by novel recurrent CHRNG mutations.J Hum Genet. 2015 Apr;60(4):213-5. doi: 10.1038/jhg.2015.2. Epub 2015 Jan 22. J Hum Genet. 2015. PMID: 25608830
References
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
