FIM, a novel FTIR-based imaging method for high throughput locomotion analysis
- PMID: 23349775
- PMCID: PMC3549958
- DOI: 10.1371/journal.pone.0053963
FIM, a novel FTIR-based imaging method for high throughput locomotion analysis
Abstract
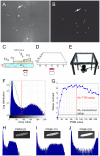
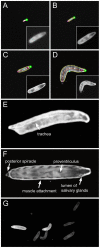
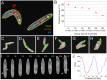
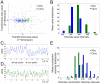
We designed a novel imaging technique based on frustrated total internal reflection (FTIR) to obtain high resolution and high contrast movies. This FTIR-based Imaging Method (FIM) is suitable for a wide range of biological applications and a wide range of organisms. It operates at all wavelengths permitting the in vivo detection of fluorescent proteins. To demonstrate the benefits of FIM, we analyzed large groups of crawling Drosophila larvae. The number of analyzable locomotion tracks was increased by implementing a new software module capable of preserving larval identity during most collision events. This module is integrated in our new tracking program named FIMTrack which subsequently extracts a number of features required for the analysis of complex locomotion phenotypes. FIM enables high throughput screening for even subtle behavioral phenotypes. We tested this newly developed setup by analyzing locomotion deficits caused by the glial knockdown of several genes. Suppression of kinesin heavy chain (khc) or rab30 function led to contraction pattern or head sweeping defects, which escaped in previous analysis. Thus, FIM permits forward genetic screens aimed to unravel the neural basis of behavior.
Conflict of interest statement
Figures
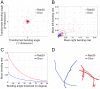
References
-
- Maimon G, Straw AD, Dickinson MH (2008) A simple vision-based algorithm for decision making in flying Drosophila. Curr Biol 18: 464–470. - PubMed
-
- Frye MA, Dickinson MH (2004) Closing the loop between neurobiology and flight behavior in Drosophila. Curr Opin Neurobiol 14: 729–736. - PubMed
-
- Fry SN, Sayaman R, Dickinson MH (2003) The aerodynamics of free-flight maneuvers in Drosophila. Science 300: 495–498. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

