Preparing synthetic biology for the world
- PMID: 23355834
- PMCID: PMC3554958
- DOI: 10.3389/fmicb.2013.00005
Preparing synthetic biology for the world
Abstract
Synthetic Biology promises low-cost, exponentially scalable products and global health solutions in the form of self-replicating organisms, or "living devices." As these promises are realized, proof-of-concept systems will gradually migrate from tightly regulated laboratory or industrial environments into private spaces as, for instance, probiotic health products, food, and even do-it-yourself bioengineered systems. What additional steps, if any, should be taken before releasing engineered self-replicating organisms into a broader user space? In this review, we explain how studies of genetically modified organisms lay groundwork for the future landscape of biosafety. Early in the design process, biological engineers are anticipating potential hazards and developing innovative tools to mitigate risk. Here, we survey lessons learned, ongoing efforts to engineer intrinsic biocontainment, and how different stakeholders in synthetic biology can act to accomplish best practices for biosafety.
Keywords: biosafety research; containment of biohazards; risk assessment; synthetic biology.
Figures

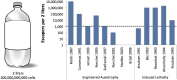

References
LinkOut - more resources
Full Text Sources
Other Literature Sources

