Detection of low-abundance KRAS mutations in colorectal cancer using microfluidic capillary electrophoresis-based restriction fragment length polymorphism method with optimized assay conditions
- PMID: 23355875
- PMCID: PMC3552804
- DOI: 10.1371/journal.pone.0054510
Detection of low-abundance KRAS mutations in colorectal cancer using microfluidic capillary electrophoresis-based restriction fragment length polymorphism method with optimized assay conditions
Abstract
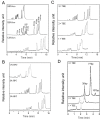
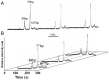
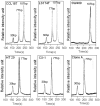
Constitutively active KRAS mutations have been found to be involved in various processes of cancer development, and render tumor cells resistant to EGFR-targeted therapies. Mutation detection methods with higher sensitivity will increase the possibility of choosing the correct individual therapy. Here, we established a highly sensitive and efficient microfluidic capillary electrophoresis-based restriction fragment length polymorphism (µCE-based RFLP) platform for low-abundance KRAS genotyping with the combination of µCE and RFLP techniques. By using our self-built sensitive laser induced fluorescence (LIF) detector and a new DNA intercalating dye YOYO-1, the separation conditions of µCE for ΦX174 HaeIII DNA marker were first optimized. Then, a Mav I digested 107-bp KRAS gene fragment was directly introduced into the microfluidic device and analyzed by µCE, in which field amplified sample stacking (FASS) technique was employed to obtain the enrichment of the RFLP digestion products and extremely improved the sensitivity. The accurate analysis of KRAS statuses in HT29, LS174T, CCL187, SW480, Clone A, and CX-1 colorectal cancer (CRC) cell lines by µCE-based RFLP were achieved in 5 min with picoliter-scale sample consumption, and as low as 0.01% of mutant KRAS could be identified from a large excess of wild-type genomic DNA (gDNA). In 98 paraffin-embedded CRC tissues, KRAS codon 12 mutations were discovered in 28 (28.6%), significantly higher than that obtained by direct sequencing (13, 13.3%). Clone sequencing confirmed these results and showed this system could detect at least 0.4% of the mutant KRAS in CRC tissue slides. Compared with direct sequencing, the new finding of the µCE-based RFLP platform was that KRAS mutations in codon 12 were correlated with the patient's age. In conclusion, we established a sensitive, fast, and cost-effective screening method for KRAS mutations, and successfully detected low-abundance KRAS mutations in clinical samples, which will allow provision of more precise individualized cancer therapy.
Conflict of interest statement
Figures



Similar articles
-
Optimization of routine KRAS mutation PCR-based testing procedure for rational individualized first-line-targeted therapy selection in metastatic colorectal cancer.Cancer Med. 2013 Feb;2(1):11-20. doi: 10.1002/cam4.47. Epub 2013 Feb 3. Cancer Med. 2013. PMID: 24133623 Free PMC article.
-
Highly sensitive detection of the PIK3CA (H1047R) mutation in colorectal cancer using a novel PCR-RFLP method.BMC Cancer. 2016 Jul 12;16:454. doi: 10.1186/s12885-016-2493-9. BMC Cancer. 2016. PMID: 27405731 Free PMC article.
-
Detection of KRAS codon 12 and 13 mutations by mutant-enriched PCR assay.Clin Chim Acta. 2014 Sep 25;436:169-75. doi: 10.1016/j.cca.2014.05.008. Epub 2014 May 24. Clin Chim Acta. 2014. PMID: 24863805
-
Recommendations from the EGAPP Working Group: can testing of tumor tissue for mutations in EGFR pathway downstream effector genes in patients with metastatic colorectal cancer improve health outcomes by guiding decisions regarding anti-EGFR therapy?Genet Med. 2013 Jul;15(7):517-27. doi: 10.1038/gim.2012.184. Epub 2013 Feb 21. Genet Med. 2013. PMID: 23429431
-
Advances in capillary electrophoresis and the implications for drug discovery.Expert Opin Drug Discov. 2017 Feb;12(2):213-224. doi: 10.1080/17460441.2017.1268121. Epub 2016 Dec 9. Expert Opin Drug Discov. 2017. PMID: 27911223 Free PMC article. Review.
Cited by
-
Lack of association between let-7 binding site polymorphism rs712 and risk of nasopharyngeal carcinoma.Fam Cancer. 2014 Mar;13(1):93-7. doi: 10.1007/s10689-013-9681-4. Fam Cancer. 2014. PMID: 23996697
-
Performance of probe polymerization-conjunction-agarose gel electrophoresis in the rapid detection of KRAS gene mutation.Genet Mol Biol. 2018 Jul/Sept.;41(3):555-561. doi: 10.1590/1678-4685-GMB-2017-0197. Epub 2018 Jul 16. Genet Mol Biol. 2018. PMID: 30080912 Free PMC article.
-
A Low-Cost Palmtop High-Speed Capillary Electrophoresis Bioanalyzer with Laser Induced Fluorescence Detection.Sci Rep. 2018 Jan 29;8(1):1791. doi: 10.1038/s41598-018-20058-0. Sci Rep. 2018. PMID: 29379053 Free PMC article.
-
Microfluidic approaches for cell-based molecular diagnosis.Biomicrofluidics. 2018 Sep 14;12(5):051501. doi: 10.1063/1.5030891. eCollection 2018 Sep. Biomicrofluidics. 2018. PMID: 30271515 Free PMC article.
-
Prevalence of K-RAS mutations and CA125 tumor marker in patients with ovarian carcinoma.Caspian J Intern Med. 2020 Winter;11(1):75-82. doi: 10.22088/cjim.11.1.75. Caspian J Intern Med. 2020. PMID: 32042390 Free PMC article.
References
-
- Jemal A, Siegel R, Ward E, Hao Y, Xu J, et al. (2009) Cancer statistics, 2009. CA Cancer J Clin 59: 225–249. - PubMed
-
- Van Cutsem E, Köhne CH, Hitre E, Zaluski J, Chang Chien CR, et al. (2009) Cetuximab and chemotherapy as initial treatment for metastatic colorectal cancer. N Engl J Med 360: 1408–1417. - PubMed
-
- Ma WW, Adjei AA (2009) Novel agents on the horizon for cancer therapy. CA Cancer J Clin 59: 111–137. - PubMed
-
- Loupakis F, Cremolini C, Salvatore L, Schirripa M, Lonardi S, et al. (2012) Clinical impact of anti-epidermal growth factor receptor monoclonal antibodies in first-line treatment of metastatic colorectal cancer: Meta-analytical estimation and implications for therapeutic strategies. Cancer 118: 1523–1532. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

