Cyr61, a matricellular protein, is needed for dendritic arborization of hippocampal neurons
- PMID: 23362279
- PMCID: PMC3605668
- DOI: 10.1074/jbc.M112.411629
Cyr61, a matricellular protein, is needed for dendritic arborization of hippocampal neurons
Abstract
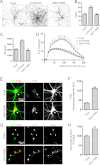
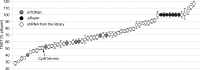
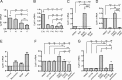
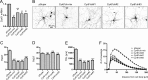
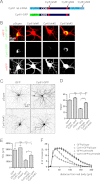
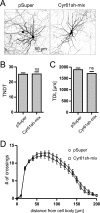
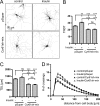
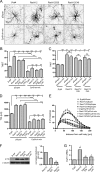
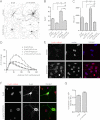
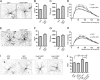
The shape of the dendritic arbor is one of the criteria of neuron classification and reflects functional specialization of particular classes of neurons. The development of a proper dendritic branching pattern strongly relies on interactions between the extracellular environment and intracellular processes responsible for dendrite growth and stability. We previously showed that mammalian target of rapamycin (mTOR) kinase is crucial for this process. In this work, we performed a screen for modifiers of dendritic growth in hippocampal neurons, the expression of which is potentially regulated by mTOR. As a result, we identified Cyr61, an angiogenic factor with unknown neuronal function, as a novel regulator of dendritic growth, which controls dendritic growth in a β1-integrin-dependent manner.
Figures
Similar articles
-
Regulation of dendritic morphogenesis by Ras-PI3K-Akt-mTOR and Ras-MAPK signaling pathways.J Neurosci. 2005 Dec 7;25(49):11288-99. doi: 10.1523/JNEUROSCI.2284-05.2005. J Neurosci. 2005. PMID: 16339024 Free PMC article.
-
Control of dendritic arborization by the phosphoinositide-3'-kinase-Akt-mammalian target of rapamycin pathway.J Neurosci. 2005 Dec 7;25(49):11300-12. doi: 10.1523/JNEUROSCI.2270-05.2005. J Neurosci. 2005. PMID: 16339025 Free PMC article.
-
Mammalian target of rapamycin complex 1 (mTORC1) and 2 (mTORC2) control the dendritic arbor morphology of hippocampal neurons.J Biol Chem. 2012 Aug 31;287(36):30240-56. doi: 10.1074/jbc.M112.374405. Epub 2012 Jul 18. J Biol Chem. 2012. PMID: 22810227 Free PMC article.
-
The integrated role of ACh, ERK and mTOR in the mechanisms of hippocampal inhibitory avoidance memory.Neurobiol Learn Mem. 2015 Mar;119:18-33. doi: 10.1016/j.nlm.2014.12.014. Epub 2015 Jan 13. Neurobiol Learn Mem. 2015. PMID: 25595880 Review.
-
Protocadherins branch out: Multiple roles in dendrite development.Cell Adh Migr. 2015;9(3):214-26. doi: 10.1080/19336918.2014.1000069. Epub 2015 Apr 14. Cell Adh Migr. 2015. PMID: 25869446 Free PMC article. Review.
Cited by
-
An Adaptive Role for DNA Double-Strand Breaks in Hippocampus-Dependent Learning and Memory.Int J Mol Sci. 2022 Jul 28;23(15):8352. doi: 10.3390/ijms23158352. Int J Mol Sci. 2022. PMID: 35955487 Free PMC article. Review.
-
ESCRT Proteins Control the Dendritic Morphology of Developing and Mature Hippocampal Neurons.Mol Neurobiol. 2019 Jul;56(7):4866-4879. doi: 10.1007/s12035-018-1418-9. Epub 2018 Nov 7. Mol Neurobiol. 2019. PMID: 30406428 Free PMC article.
-
GSK3α and GSK3β Phosphorylate Arc and Regulate its Degradation.Front Mol Neurosci. 2017 Jun 16;10:192. doi: 10.3389/fnmol.2017.00192. eCollection 2017. Front Mol Neurosci. 2017. PMID: 28670266 Free PMC article.
-
Matricellular proteins of the Cyr61/CTGF/NOV (CCN) family and the nervous system.Front Cell Neurosci. 2015 Jun 24;9:237. doi: 10.3389/fncel.2015.00237. eCollection 2015. Front Cell Neurosci. 2015. PMID: 26157362 Free PMC article. Review.
-
Cyr61 promotes Schwann cell proliferation and migration via αvβ3 integrin.BMC Mol Cell Biol. 2021 Apr 7;22(1):21. doi: 10.1186/s12860-021-00360-y. BMC Mol Cell Biol. 2021. PMID: 33827416 Free PMC article.
References
-
- Stuart G., Spruston N., Häusser M. (2007) Dendrites, 2nd Ed., Oxford University Press, Oxford
-
- Segev I., London M. (2000) Untangling dendrites with quantitative models. Science 290, 744–750 - PubMed
-
- Urbanska M., Blazejczyk M., Jaworski J. (2008) Molecular basis of dendritic arborization. Acta Neurobiol. Exp. 68, 264–288 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

