Metagenomic profile of gut microbiota in children during cholera and recovery
- PMID: 23369162
- PMCID: PMC3574833
- DOI: 10.1186/1757-4749-5-1
Metagenomic profile of gut microbiota in children during cholera and recovery
Abstract
Background: The diverse bacterial communities colonizing the gut (gastrointestinal tract) of infants as commensal flora, which play an important role in nutrient absorption and determining the state of health, are known to alter due to diarrhea.
Method: Bacterial community dynamics in children suffering from cholera and during recovery period were examined in the present study by employing metagenomic tool, followed by DNA sequencing and analysis. For this, bacterial community DNA was extracted from fecal samples of nine clinically confirmed cholera children (age 2-3 years) at day 0 (acute cholera), day 2 (antibiotic therapy), day 7 and, and day 28, and the variable region of 16S rRNA genes were amplified by universal primer PCR.
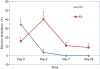
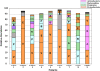
Results: 454 parallel sequencing of the amplified DNA followed by similarity search of the sequenced data against an rRNA database allowed us to identify V. cholerae, the cause of cholera, in all nine children at day 0, and as predominant species in six children, accounting for 35% of the total gut microbiota on an average in all the nine children. The relative abundance (mean ± sem %) of bacteria belonging to phyla Proteobacteria, Firmicutes, Bacteroidetes, and Actinobacteria, was 55 ± 7, 18 ± 4, 13 ± 4, and 8 ± 4, respectively, at day 0, while these values were 12 ± 4, 43 ± 4, 33 ± 3, and 12 ± 2, respectively, at day 28. As antibiotic therapy began, V. cholerae count declined significantly (p< 0.001) and was found only in four children at day 2 and two children at day 7 with the relative abundance of 3.7% and 0.01%, respectively, which continued up to day 28 in the two children. Compared to acute cholera condition (day 0), the relative abundance of Escherichia coli, Enterococcus, and Veillonella increased at day 2 (antibiotic therapy) while Bifidobacterium, Bacteroides, and Ruminococcus decreased.
Conclusion: Cholera results expulsion of major commensal bacteria of phyla Bacteroidetes, Firmicutes, and Actinobacteria, and increase of harmful Proteobacteria to colonize the gut during acute and convalescence states. The observed microbiota disruption might explain the prevalent malnutrition in children of Bangladesh where diarrheal diseases are endemic.
Figures



Similar articles
-
Gut microbiota of healthy and malnourished children in bangladesh.Front Microbiol. 2011 Nov 21;2:228. doi: 10.3389/fmicb.2011.00228. eCollection 2011. Front Microbiol. 2011. PMID: 22125551 Free PMC article.
-
Gut microbiota shifts favorably with delivery of handwashing with soap and water treatment intervention in a prospective cohort (CHoBI7 trial).J Health Popul Nutr. 2023 Dec 21;42(1):146. doi: 10.1186/s41043-023-00477-0. J Health Popul Nutr. 2023. PMID: 38129922 Free PMC article. Clinical Trial.
-
Molecular Analysis of Selected Resistance Determinants in Diarrheal Fecal Samples Collected From Kolkata, India Reveals an Abundance of Resistance Genes and the Potential Role of the Microbiota in Its Dissemination.Front Public Health. 2020 Mar 11;8:61. doi: 10.3389/fpubh.2020.00061. eCollection 2020. Front Public Health. 2020. PMID: 32219088 Free PMC article.
-
Gallstone Disease, Obesity and the Firmicutes/Bacteroidetes Ratio as a Possible Biomarker of Gut Dysbiosis.J Pers Med. 2020 Dec 25;11(1):13. doi: 10.3390/jpm11010013. J Pers Med. 2020. PMID: 33375615 Free PMC article. Review.
-
Metagenomic insights into the roles of Proteobacteria in the gastrointestinal microbiomes of healthy dogs and cats.Microbiologyopen. 2018 Oct;7(5):e00677. doi: 10.1002/mbo3.677. Epub 2018 Jun 17. Microbiologyopen. 2018. PMID: 29911322 Free PMC article. Review.
Cited by
-
Gut microbial succession follows acute secretory diarrhea in humans.mBio. 2015 May 19;6(3):e00381-15. doi: 10.1128/mBio.00381-15. mBio. 2015. PMID: 25991682 Free PMC article.
-
Rotavirus infection induces glycan availability to promote ileum-specific changes in the microbiome aiding rotavirus virulence.Gut Microbes. 2020 Sep 2;11(5):1324-1347. doi: 10.1080/19490976.2020.1754714. Epub 2020 May 13. Gut Microbes. 2020. PMID: 32404017 Free PMC article.
-
Changing facades of Vibrio cholerae: An enigma in the epidemiology of cholera.Indian J Med Res. 2018 Feb;147(2):133-141. doi: 10.4103/ijmr.IJMR_280_17. Indian J Med Res. 2018. PMID: 29806601 Free PMC article. Review.
-
Vibrio cholerae at the Intersection of Immunity and the Microbiome.mSphere. 2019 Nov 27;4(6):e00597-19. doi: 10.1128/mSphere.00597-19. mSphere. 2019. PMID: 31776240 Free PMC article. Review.
-
Microbiome Profiling of Enterotoxigenic Escherichia coli (ETEC) Carriers Highlights Signature Differences between Symptomatic and Asymptomatic Individuals.mBio. 2022 Jun 28;13(3):e0015722. doi: 10.1128/mbio.00157-22. Epub 2022 May 10. mBio. 2022. PMID: 35536001 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

