Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging
- PMID: 23371551
- PMCID: PMC3916822
- DOI: 10.1126/science.1230593
Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging
Abstract
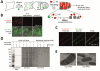
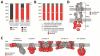
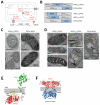
Microscopy and mass spectrometry (MS) are complementary techniques: The former provides spatiotemporal information in living cells, but only for a handful of recombinant proteins at a time, whereas the latter can detect thousands of endogenous proteins simultaneously, but only in lysed samples. Here, we introduce technology that combines these strengths by offering spatially and temporally resolved proteomic maps of endogenous proteins within living cells. Our method relies on a genetically targetable peroxidase enzyme that biotinylates nearby proteins, which are subsequently purified and identified by MS. We used this approach to identify 495 proteins within the human mitochondrial matrix, including 31 not previously linked to mitochondria. The labeling was exceptionally specific and distinguished between inner membrane proteins facing the matrix versus the intermembrane space (IMS). Several proteins previously thought to reside in the IMS or outer membrane, including protoporphyrinogen oxidase, were reassigned to the matrix by our proteomic data and confirmed by electron microscopy. The specificity of peroxidase-mediated proteomic mapping in live cells, combined with its ease of use, offers biologists a powerful tool for understanding the molecular composition of living cells.
Figures
Comment in
-
Proteomics: Marked for depth.Nat Methods. 2013 Apr;10(4):284. doi: 10.1038/nmeth.2427. Nat Methods. 2013. PMID: 23653924 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

