Arabidopsis paired amphipathic helix proteins SNL1 and SNL2 redundantly regulate primary seed dormancy via abscisic acid-ethylene antagonism mediated by histone deacetylation
- PMID: 23371947
- PMCID: PMC3584531
- DOI: 10.1105/tpc.112.108191
Arabidopsis paired amphipathic helix proteins SNL1 and SNL2 redundantly regulate primary seed dormancy via abscisic acid-ethylene antagonism mediated by histone deacetylation
Abstract
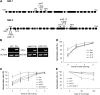
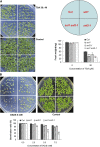
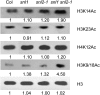
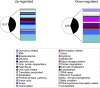
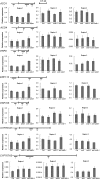
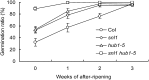
Histone (de)acetylation is a highly conserved chromatin modification that is vital for development and growth. In this study, we identified a role in seed dormancy for two members of the histone deacetylation complex in Arabidopsis thaliana, SIN3-LIKE1 (SNL1) and SNL2. The double mutant snl1 snl2 shows reduced dormancy and hypersensitivity to the histone deacetylase inhibitors trichostatin A and diallyl disulfide compared with the wild type. SNL1 interacts with HISTONE DEACETYLASE19 in vitro and in planta, and loss-of-function mutants of SNL1 and SNL2 show increased acetylation levels of histone 3 lysine 9/18 (H3K9/18) and H3K14. Moreover, SNL1 and SNL2 regulate key genes involved in the ethylene and abscisic acid (ABA) pathways by decreasing their histone acetylation levels. Taken together, we showed that SNL1 and SNL2 regulate seed dormancy by mediating the ABA-ethylene antagonism in Arabidopsis. SNL1 and SNL2 could represent a cross-link point of the ABA and ethylene pathways in the regulation of seed dormancy.
Figures





Similar articles
-
Arabidopsis seed germination speed is controlled by SNL histone deacetylase-binding factor-mediated regulation of AUX1.Nat Commun. 2016 Nov 11;7:13412. doi: 10.1038/ncomms13412. Nat Commun. 2016. PMID: 27834370 Free PMC article.
-
HISTONE DEACETYLASE19 interacts with HSL1 and participates in the repression of seed maturation genes in Arabidopsis seedlings.Plant Cell. 2013 Jan;25(1):134-48. doi: 10.1105/tpc.112.096313. Epub 2013 Jan 29. Plant Cell. 2013. PMID: 23362207 Free PMC article.
-
AtPER1 enhances primary seed dormancy and reduces seed germination by suppressing the ABA catabolism and GA biosynthesis in Arabidopsis seeds.Plant J. 2020 Jan;101(2):310-323. doi: 10.1111/tpj.14542. Epub 2019 Oct 22. Plant J. 2020. PMID: 31536657
-
Regulation of seed dormancy by histone post-translational modifications in the model plant Arabidopsis thaliana.J Exp Bot. 2024 Oct 16;75(19):6159-6166. doi: 10.1093/jxb/erae236. J Exp Bot. 2024. PMID: 38769701 Review.
-
Histone modifications affecting plant dormancy and dormancy release: common regulatory effects on hormone metabolism.J Exp Bot. 2024 Oct 16;75(19):6142-6158. doi: 10.1093/jxb/erae205. J Exp Bot. 2024. PMID: 38721634 Review.
Cited by
-
A New Intra-Specific and High-Resolution Genetic Map of Eggplant Based on a RIL Population, and Location of QTLs Related to Plant Anthocyanin Pigmentation and Seed Vigour.Genes (Basel). 2020 Jul 4;11(7):745. doi: 10.3390/genes11070745. Genes (Basel). 2020. PMID: 32635424 Free PMC article.
-
Seed dormancy and germination-emerging mechanisms and new hypotheses.Front Plant Sci. 2014 May 28;5:233. doi: 10.3389/fpls.2014.00233. eCollection 2014. Front Plant Sci. 2014. PMID: 24904627 Free PMC article. Review.
-
Transcriptome and Degradome Sequencing Reveals Dormancy Mechanisms of Cunninghamia lanceolata Seeds.Plant Physiol. 2016 Dec;172(4):2347-2362. doi: 10.1104/pp.16.00384. Epub 2016 Oct 19. Plant Physiol. 2016. PMID: 27760880 Free PMC article.
-
Research Progress on the Mechanism and Function of Histone Acetylation Regulating the Interaction between Pathogenic Fungi and Plant Hosts.J Fungi (Basel). 2024 Jul 26;10(8):522. doi: 10.3390/jof10080522. J Fungi (Basel). 2024. PMID: 39194848 Free PMC article. Review.
-
Seed Transcriptome Annotation Reveals Enhanced Expression of Genes Related to ROS Homeostasis and Ethylene Metabolism at Alternating Temperatures in Wild Cardoon.Plants (Basel). 2020 Sep 18;9(9):1225. doi: 10.3390/plants9091225. Plants (Basel). 2020. PMID: 32961840 Free PMC article.
References
-
- Audic S., Claverie J.M. (1997). The significance of digital gene expression profiles. Genome Res. 7: 986–995 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

