The circadian epigenome: how metabolism talks to chromatin remodeling
- PMID: 23385084
- PMCID: PMC4573393
- DOI: 10.1016/j.ceb.2013.01.003
The circadian epigenome: how metabolism talks to chromatin remodeling
Abstract
Circadian rhythms occur in most of the living organisms, and with a 24 hour periodicity govern a number of physiological and metabolic functions. During the past few years, an important research effort has uncovered new trails that intersect between circadian rhythms and metabolic pathways. At a molecular level, the clock machinery is responsible for the establishment of a circadian epigenome, and this can be modulated by metabolic cues. Indeed, metabolic control by the circadian clock is manifest in the development of metabolic diseases when circadian rhythms are impaired. Thus, pharmacological modulation of circadian rhythms promises new avenues for the treatment of metabolic and sleep disorders.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures


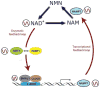
References
-
- Schibler U, Sassone-Corsi P. A web of circadian pacemakers. Cell. 2002;111(7):919–922. - PubMed
-
- Ralph MR, Foster RG, Davis FC, Menaker M. Transplanted suprachiasmatic nucleus determines circadian period. Science. 1990;247(4945):975–978. - PubMed
-
- Freedman MS, Lucas RJ, Soni B, von Schantz M, Munoz M, David-Gray Z, Foster R. Regulation of mammalian circadian behavior by non-rod, non-cone, ocular photoreceptors. Science. 1999;284(5413):502–504. - PubMed
-
- Cermakian N, Sassone-Corsi P. Environmental stimulus perception and control of circadian clocks. Curr Opin Neurobiol. 2002;12(4):359–365. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

