The microtubule-binding protein ensconsin is an essential cofactor of kinesin-1
- PMID: 23394833
- PMCID: PMC3580027
- DOI: 10.1016/j.cub.2013.01.008
The microtubule-binding protein ensconsin is an essential cofactor of kinesin-1
Abstract
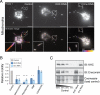
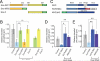
Kinesin-1 is a major microtubule motor that drives transport of numerous cellular cargoes toward the plus ends of microtubules. In the cell, kinesin-1 exists primarily in an inactive, autoinhibited state, and motor activation is thought to occur upon binding to cargo through the C terminus. Using RNAi-mediated depletion in Drosophila S2 cells, we demonstrate that kinesin-1 requires ensconsin (MAP7, E-MAP-115), a ubiquitous microtubule-associated protein, for its primary function of organelle transport. We show that ensconsin is required for organelle transport in Drosophila neurons and that Drosophila homozygous for ensconsin gene deletion are unable to survive to adulthood. An ensconsin N-terminal truncation that cannot bind microtubules is sufficient to activate organelle transport by kinesin-1, indicating that this activating domain functions independently of microtubule binding. Interestingly, ens mutant flies retaining expression of this truncation show normal viability. A "hingeless" mutant of kinesin-1, which mimics the active conformation of the motor, does not require ensconsin for transport in S2 cells, suggesting that ensconsin plays a role in relieving autoinhibition of kinesin-1. Together with other recent work, our study suggests that ensconsin is an essential cofactor for all known functions of kinesin-1.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures


References
-
- Friedman DS, Vale RD. Single-molecule analysis of kinesin motility reveals regulation by the cargo-binding tail domain. Nat Cell Biol. 1999;1:293–297. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

