A genomic signature and the identification of new sporulation genes
- PMID: 23396918
- PMCID: PMC3624599
- DOI: 10.1128/JB.02110-12
A genomic signature and the identification of new sporulation genes
Abstract
Bacterial endospores are the most resistant cell type known to humans, as they are able to withstand extremes of temperature, pressure, chemical injury, and time. They are also of interest because the endospore is the infective particle in a variety of human and livestock diseases. Endosporulation is characterized by the morphogenesis of an endospore within a mother cell. Based on the genes known to be involved in endosporulation in the model organism Bacillus subtilis, a conserved core of about 100 genes was derived, representing the minimal machinery for endosporulation. The core was used to define a genomic signature of about 50 genes that are able to distinguish endospore-forming organisms, based on complete genome sequences, and we show this 50-gene signature is robust against phylogenetic proximity and other artifacts. This signature includes previously uncharacterized genes that we can now show are important for sporulation in B. subtilis and/or are under developmental control, thus further validating this genomic signature. We also predict that a series of polyextremophylic organisms, as well as several gut bacteria, are able to form endospores, and we identified 3 new loci essential for sporulation in B. subtilis: ytaF, ylmC, and ylzA. In all, the results support the view that endosporulation likely evolved once, at the base of the Firmicutes phylum, and is unrelated to other bacterial cell differentiation programs and that this involved the evolution of new genes and functions, as well as the cooption of ancestral, housekeeping functions.
Figures



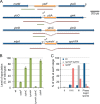

References
-
- Nicholson WL. 2004. Ubiquity, longevity, and ecological roles of Bacillus spores, p 1–15 In Ricca E, Henriques AO, Cutting SM. (ed), Bacterial spore formers: probiotics and emerging applications. Horizon Scientific Press, London, England
-
- Setlow P. 2006. Spores of Bacillus subtilis: their resistance to and killing by radiation, heat and chemicals. J. Appl. Microbiol. 101:514–525 - PubMed
-
- Stenfors Arnesen LP, Fagerlund A, Granum PE. 2008. From soil to gut: Bacillus cereus and its food poisoning toxins. FEMS Microbiol. Rev. 32:579–606 - PubMed
-
- Fouet A, Mock M. 2006. Regulatory networks for virulence and persistence of Bacillus anthracis. Curr. Opin. Microbiol. 9:160–166 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

