Persistent whole-chromosome aneuploidy is generally associated with nascent allohexaploid wheat
- PMID: 23401544
- PMCID: PMC3587266
- DOI: 10.1073/pnas.1300153110
Persistent whole-chromosome aneuploidy is generally associated with nascent allohexaploid wheat
Abstract
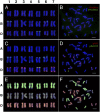
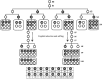
Allopolyploidization has been a driving force in plant evolution. Formation of common wheat (Triticum aestivum L.) represents a classic example of successful speciation via allopolyploidy. Nevertheless, the immediate chromosomal consequences of allopolyploidization in wheat remain largely unexplored. We report here an in-depth investigation on transgenerational chromosomal variation in resynthesized allohexaploid wheats that are identical in genome constitution to common wheat. We deployed sequential FISH, genomic in situ hybridization (GISH), and homeolog-specific pyrosequencing, which enabled unequivocal identification of each of the 21 homologous chromosome pairs in each of >1,000 individual plants from 16 independent lines. We report that whole-chromosome aneuploidy occurred ubiquitously in early generations (from selfed generation S(1) to >S(20)) of wheat allohexaploidy although at highly variable frequencies (20-100%). In contrast, other types of gross structural variations were scant. Aneuploidy included an unexpected hidden type, which had a euploid chromosome number of 2n = 42 but with simultaneous loss and gain of nonhomeologous chromosomes. Of the three constituent subgenomes, B showed the most lability for aneuploidy, followed by A, but the recently added D subgenome was largely stable in most of the studied lines. Chromosome loss and gain were also unequal across the 21 homologous chromosome pairs. Pedigree analysis showed no evidence for progressive karyotype stabilization even with multigenerational selection for euploidy. Profiling of two traits directly related to reproductive fitness showed that although pollen viability was generally reduced by aneuploidy, the adverse effect of aneuploidy on seed-set is dependent on both aneuploidy type and synthetic line.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Intrinsic karyotype stability and gene copy number variations may have laid the foundation for tetraploid wheat formation.Proc Natl Acad Sci U S A. 2013 Nov 26;110(48):19466-71. doi: 10.1073/pnas.1319598110. Epub 2013 Nov 11. Proc Natl Acad Sci U S A. 2013. PMID: 24218593 Free PMC article.
-
Global transgenerational gene expression dynamics in two newly synthesized allohexaploid wheat (Triticum aestivum) lines.BMC Biol. 2012 Jan 26;10:3. doi: 10.1186/1741-7007-10-3. BMC Biol. 2012. PMID: 22277161 Free PMC article.
-
Chromosomal and genome-wide molecular changes associated with initial stages of allohexaploidization in wheat can be transit and incidental.Genome. 2011 Aug;54(8):692-9. doi: 10.1139/g11-028. Epub 2011 Jul 29. Genome. 2011. PMID: 21797821
-
Making the Bread: Insights from Newly Synthesized Allohexaploid Wheat.Mol Plant. 2015 Jun;8(6):847-59. doi: 10.1016/j.molp.2015.02.016. Epub 2015 Mar 5. Mol Plant. 2015. PMID: 25747845 Review.
-
Allopolyploidy--a shaping force in the evolution of wheat genomes.Cytogenet Genome Res. 2005;109(1-3):250-8. doi: 10.1159/000082407. Cytogenet Genome Res. 2005. PMID: 15753584 Review.
Cited by
-
Intra- and Inter-Specific Crosses among Centaurea aspera L. (Asteraceae) Polyploid Relatives-Influences on Distribution and Polyploid Establishment.Plants (Basel). 2020 Sep 3;9(9):1142. doi: 10.3390/plants9091142. Plants (Basel). 2020. PMID: 32899362 Free PMC article.
-
Targeting Induced Local Lesions in the Wheat DEMETER and DRE2 Genes, Responsible for Transcriptional Derepression of Wheat Gluten Proteins in the Developing Endosperm.Front Nutr. 2022 Mar 3;9:847635. doi: 10.3389/fnut.2022.847635. eCollection 2022. Front Nutr. 2022. PMID: 35308262 Free PMC article.
-
Intrinsic karyotype stability and gene copy number variations may have laid the foundation for tetraploid wheat formation.Proc Natl Acad Sci U S A. 2013 Nov 26;110(48):19466-71. doi: 10.1073/pnas.1319598110. Epub 2013 Nov 11. Proc Natl Acad Sci U S A. 2013. PMID: 24218593 Free PMC article.
-
Precisely mapping a major QTL for grain weight on chromosome 5B of the founder parent Chuanmai42 in the wheat-growing region of southwestern China.Theor Appl Genet. 2023 May 31;136(6):146. doi: 10.1007/s00122-023-04383-1. Theor Appl Genet. 2023. PMID: 37258797
-
Wheat Landrace Genome Diversity.Genetics. 2017 Apr;205(4):1657-1676. doi: 10.1534/genetics.116.194688. Epub 2017 Feb 17. Genetics. 2017. PMID: 28213475 Free PMC article.
References
-
- Feldman M, Lupton FGH, Miller T. Wheats. In: Smartt J, Simmonds NW, editors. Evolution of Crops. 2nd Ed. London: Longman Scientific; 1995. pp. 184–192.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

