Transcriptomic analysis of differentially expressed genes during anther development in genetic male sterile and wild type cotton by digital gene-expression profiling
- PMID: 23402279
- PMCID: PMC3599889
- DOI: 10.1186/1471-2164-14-97
Transcriptomic analysis of differentially expressed genes during anther development in genetic male sterile and wild type cotton by digital gene-expression profiling
Abstract
Background: Cotton (Gossypium hirsutum) anther development involves a diverse range of gene interactions between sporophytic and gametophytic tissues. However, only a small number of genes are known to be specifically involved in this developmental process and the molecular mechanism of the genetic male sterility (GMS) is still poorly understand. To fully explore the global gene expression during cotton anther development and identify genes related to male sterility, a digital gene expression (DGE) analysis was adopted.
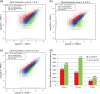
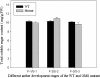
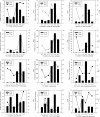
Results: Six DGE libraries were constructed from the cotton anthers of the wild type (WT) and GMS mutant (in the WT background) in three stages of anther development, resulting in 21,503 to 37,352 genes detected in WT and GMS mutant anthers. Compared with the fertile isogenic WT, 9,595 (30% of the expressed genes), 10,407 (25%), and 3,139 (10%) genes were differentially expressed at the meiosis, tetrad, and uninucleate microspore stages of GMS mutant anthers, respectively. Using both DGE experiments and real-time quantitative RT-PCR, the expression of many key genes required for anther development were suppressed in the meiosis stage and the uninucleate microspore stage in anthers of the mutant, but these genes were activated in the tetrad stage of anthers in the mutant. These genes were associated predominantly with hormone synthesis, sucrose and starch metabolism, the pentose phosphate pathway, glycolysis, flavonoid metabolism, and histone protein synthesis. In addition, several genes that participate in DNA methylation, cell wall loosening, programmed cell death, and reactive oxygen species generation/scavenging were activated during the three anther developmental stages in the mutant.
Conclusions: Compared to the same anther developmental stage of the WT, many key genes involved in various aspects of anther development show a reverse gene expression pattern in the GMS mutant, which indicates that diverse gene regulation pathways are involved in the GMS mutant anther development. These findings provide the first insights into the mechanism that leads to genetic male sterility in cotton and contributes to a better understanding of the regulatory network involved in anther development in cotton.
Figures



References
-
- Schnable PS, Wise RP. The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci. 1998;3:175–180. doi: 10.1016/S1360-1385(98)01235-7. - DOI
-
- Sheng TZ. Thesises on male sterile of cotton. Chengdu: Sichuan Science and Technology Press; 1989.
-
- Pacini E, Franchi GG, Hesse M. The tapetum: its form, function and possible phylogeny in Embryophyta. Plant Syst Evol. 1985;149:155–185. doi: 10.1007/BF00983304. - DOI
-
- Kaul ML. Male sterility in higher plants: Monograph on Theoretical and Applied Genetics. Berlin: ; 1988.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

